Publications
A near-complete list of publications is available from my Google Scholar page.
Selected manuscripts
Selected manuscripts illustrating past and current work, all with either a first or senior authorship from the Purcell lab, although much of this work was produced within the context of large consortia involving many collaborators.

Diverse NREM sleep alterations in SCZ
A spectrum of altered non-rapid eye movement sleep in schizophrenia
Kozhemiako et al. (SLEEP, 2024) [PDF]
In our second wave of data from the GRINS project, we present results that point to a spectrum of N2 sleep deficits among SCZ patients that can be measured objectively and at scale, with relevance to both the etiological heterogeneity of SCZ as well as potential iatrogenic effects of antipsychotic medication.
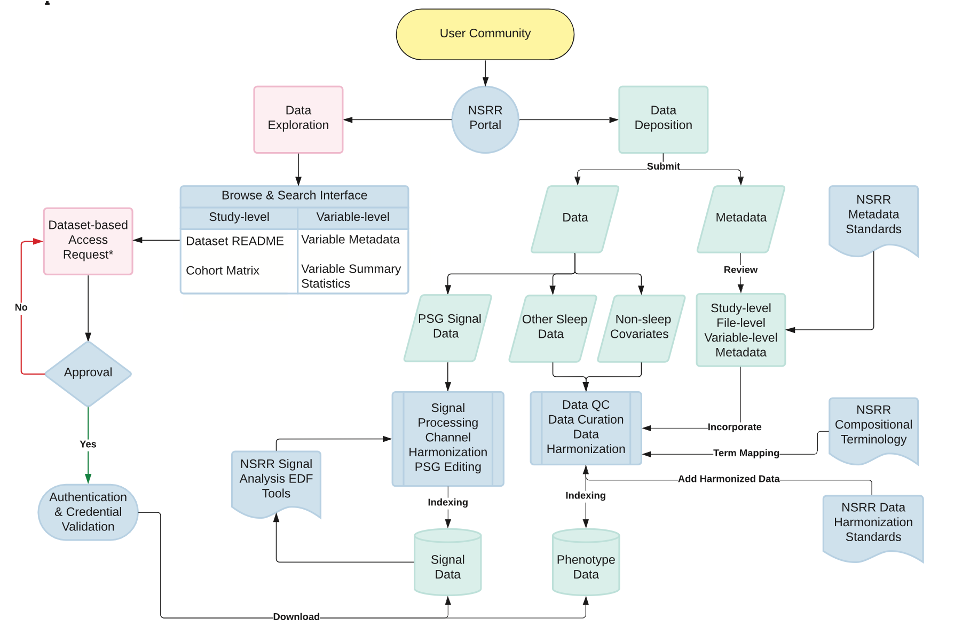
The National Sleep Research Resource: making data findable, accessible, interoperable, reusable and promoting sleep science
Zhang et al. (SLEEP, 2023) [PDF]
This paper presents a comprehensive overview of the National Sleep Research Resource (NSRR), a National Heart Lung and Blood Institute-supported repository developed to share data from clinical studies focused on the evaluation of sleep disorders.

Sleep EEG microstates and schizophrenia
Electroencephalographic Microstates During Sleep and Wake in Schizophrenia
Murphy et al. (Biological Psychiatry (GOS), 2024) [PDF]
We studied microstates during sleep in patients with schizophrenia by analyzing high-density EEG sleep data from 114 patients with schizophrenia and 79 control participants. Our findings reveal behavioral state–dependent patterns of cortical dysconnectivity in schizophrenia. Furthermore, these findings are largely unrelated to previous sleep-related EEG markers of schizophrenia such as decreased sleep spindles.

OSA and cognition
Sleep Architecture, Obstructive Sleep Apnea, and Cognitive Function in Adults
Pase et al. (JAMA Network Open, 2023) [PDF]
This study found that better sleep consolidation and the absence of OSA were associated with better global cognition over 5 years of follow-up. These findings suggest that the role of interventions to improve sleep for maintaining cognitive function requires investigation.
Sleep neurophysiology in childhood development
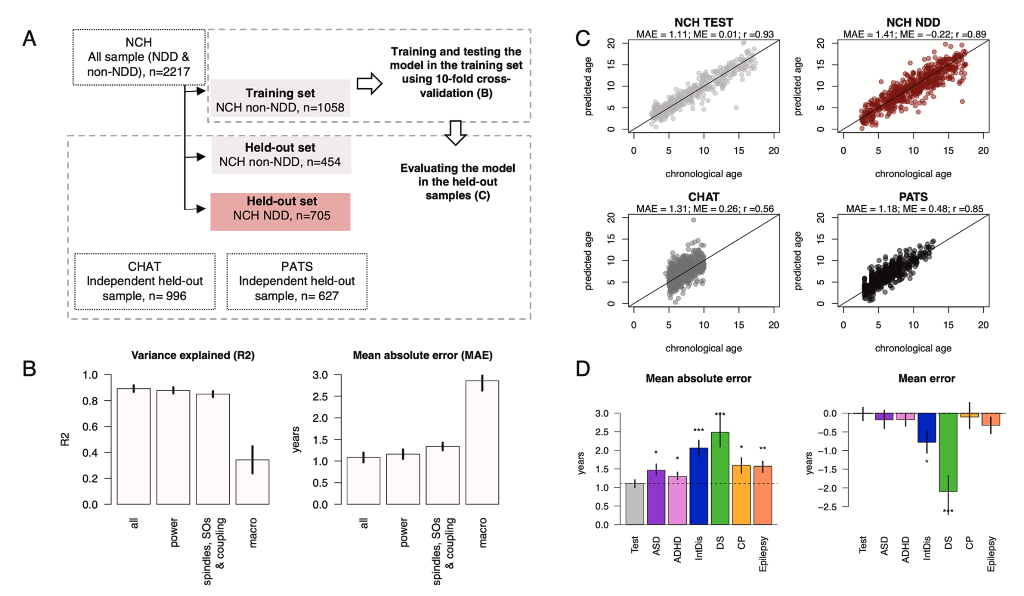
Mapping neurodevelopment with sleep macro- and micro-architecture across multiple pediatric populations
Kozhemiako et al. (NeuroImage: Clincal, 2023) [PDF]
Profiles of sleep duration and timing and corresponding EEG activity reflect brain changes that support cognitive and behavioral maturation and may provide practical markers for tracking typical and atypical neurodevelopment. To build and evaluate a sleep-based, quantitative metric of brain maturation, we used PSG data from large research and clinical samples spanning childhood and adolescence (N = 4,013, aged 2.5 to 17.5 years). Our results indicate that sleep architecture offers a sensitive window for characterizing brain maturation, suggesting the po- tential for scalable, objective sleep-based biomarkers to measure neurodevelopment.
New algorithms for EEG microstate testing
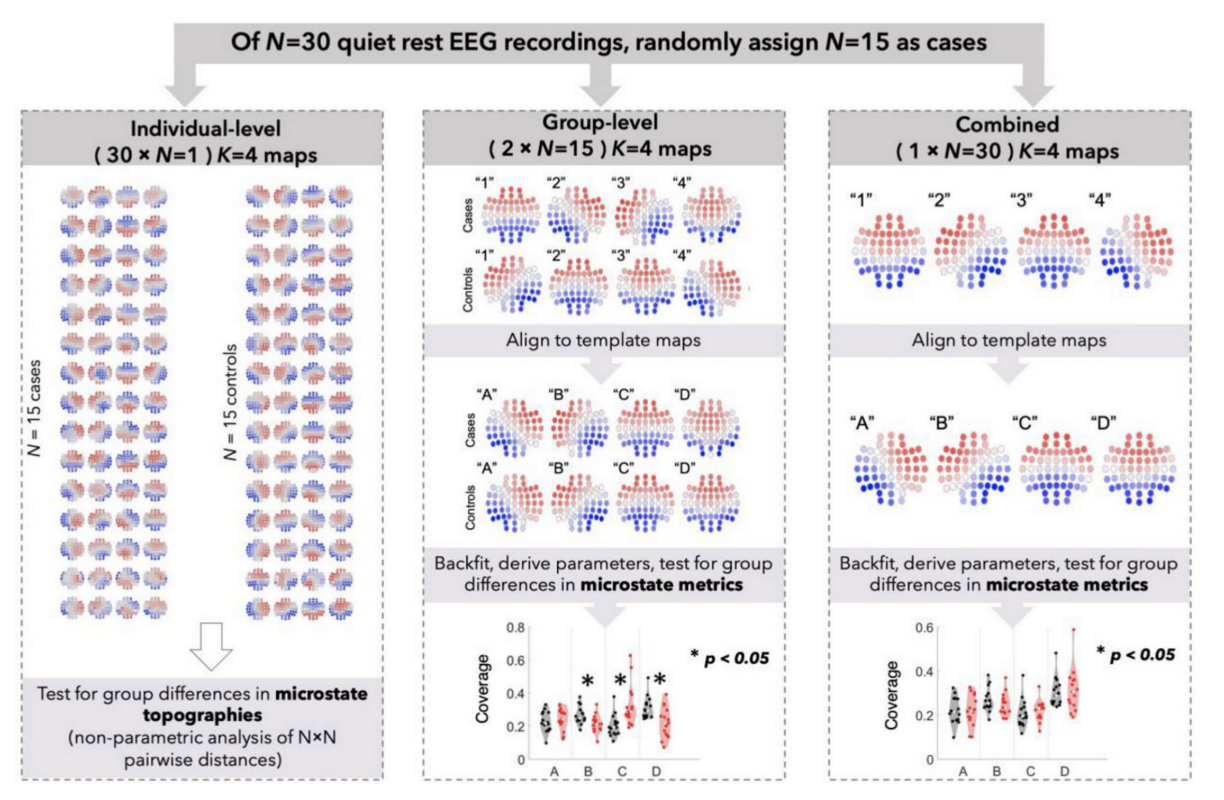
A Potential Source of Bias in Group‑Level EEG Microstate Analysis
Murphy et al. (Brain Topography, 2023) [PDF]
Microstate analysis is a promising technique for analyzing high-density electroencephalographic data, but there are multi- ple questions about methodological best practices. Our results suggest that even subtle chance differences in microstate topography can have profound effects on derived microstate metrics and that future studies using microstate analysis should take steps to mitigate this large source of error.

Tau burden and sleep architecture
Amyloid beta-independent sleep markers associated with early regional tau burden and cortical thinning
Stankeviciute et al. (Alzheimers Dement (Amst), 2023) [PDF]
In cognitively unimpaired older adults, lower SWS and higher N1 sleep were associated with higher tau burden and lower cortical thickness in brain regions associated with early tau deposition and vulnerable to AD-related neurodegeneration through mechanisms dissociable from amyloid deposition.
Characterizing sleep's EEG spectral slope
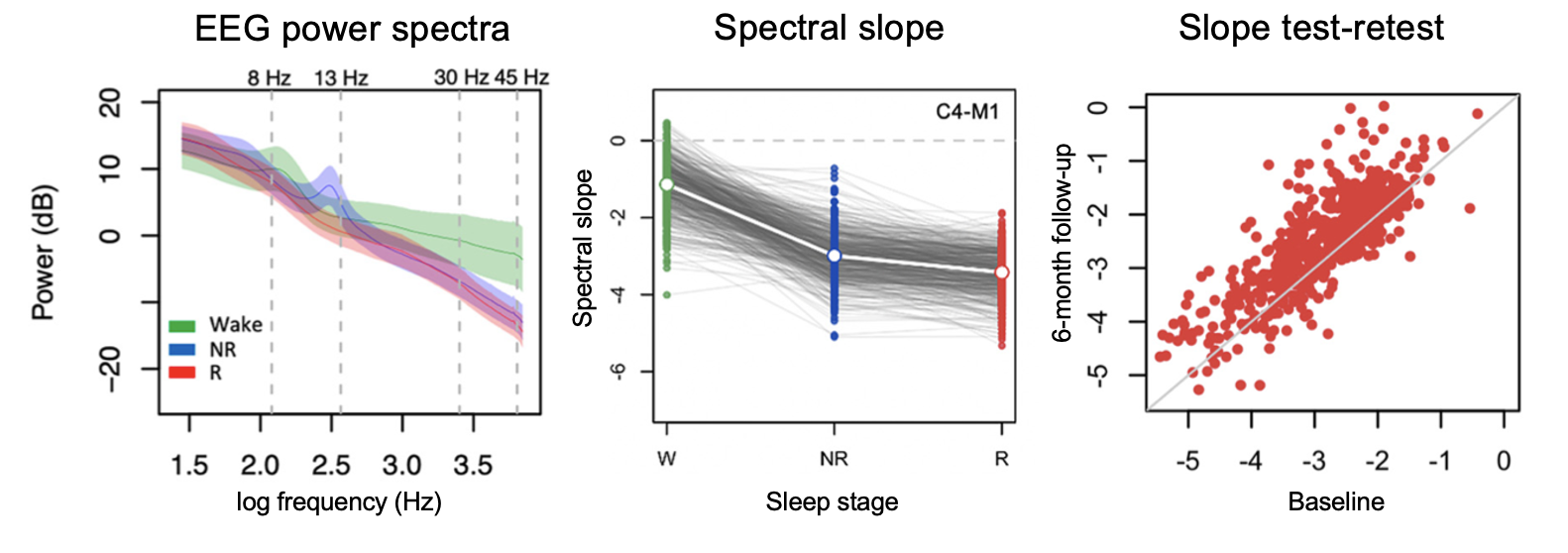
Sources of Variation in the Spectral Slope of the Sleep EEG
Kozhemiako et al. (eNeuro, 2022) [PDF]
In over 10,000 NSRR PSGs we we validate the EEG spectral slope as a neurobiological marker of cortical arousal across wake, NREM and REM sleep, and report sources of intra-individual and inter-individual variability, including changes across the lifespan.
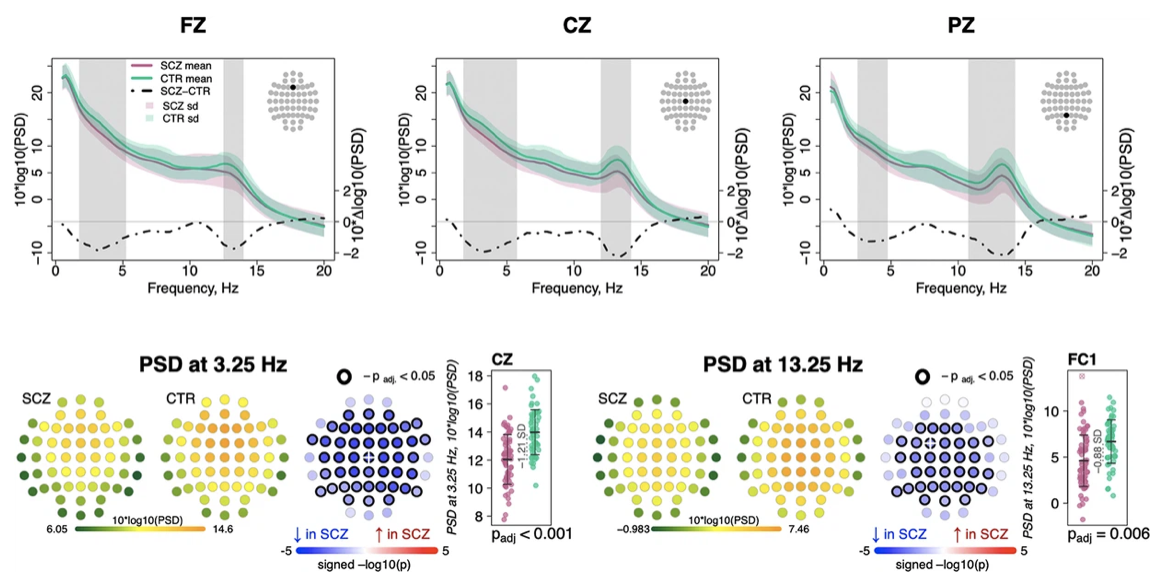
NREM spectral biomarkers of schizophrenia

Linking sleep and cognition in older adults
Macro and micro sleep architecture and cognitive performance in older adults
Djonlagic et al. (Nature Human Behaviour, 2021) [PubMed | PDF ]
This study pointed to multiple facets of sleep neurophysiology that track coherently with underlying, age-dependent determinants of cognitive and physical health trajectories in older adults.
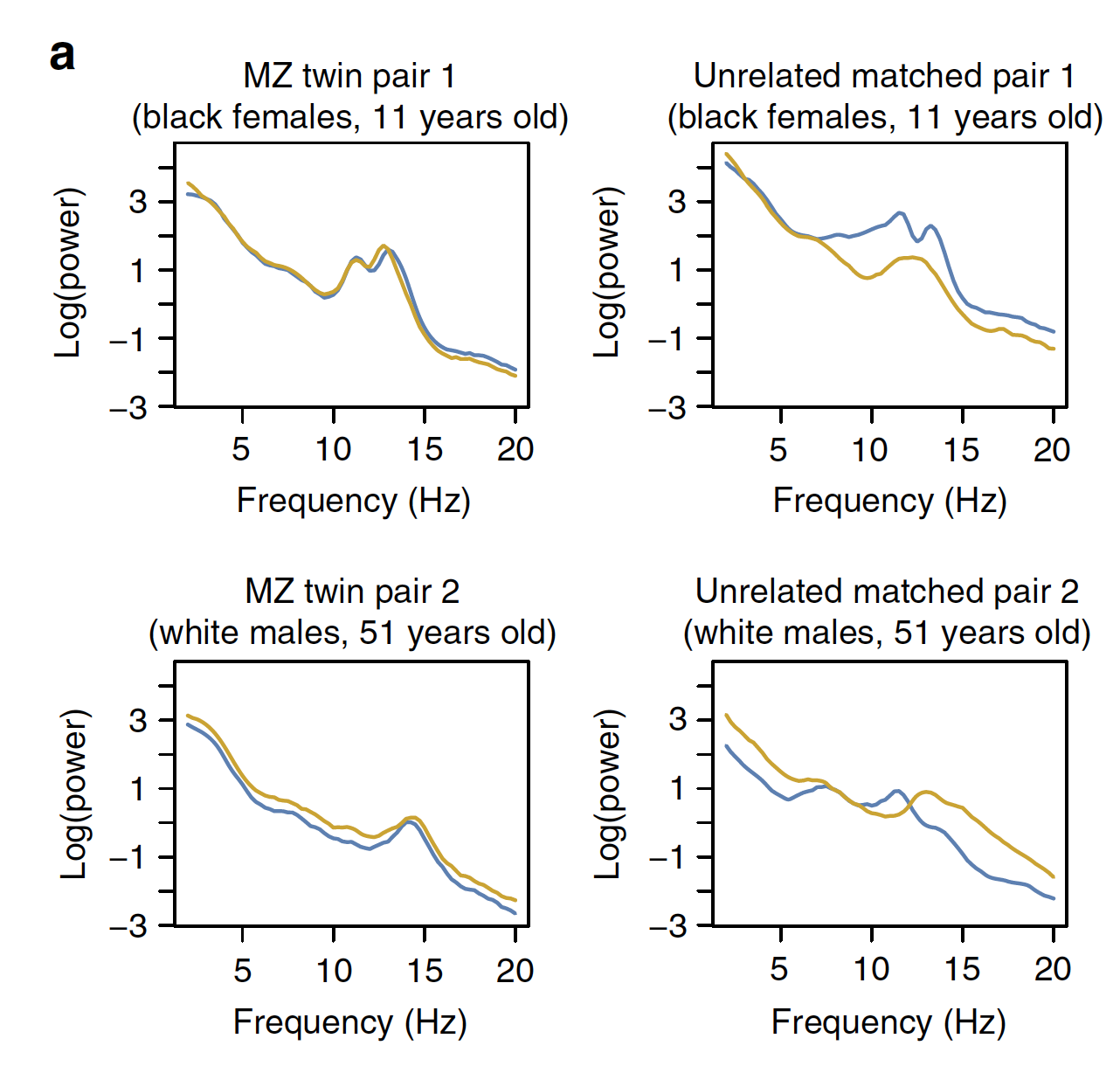
Illustrative NREM sleep power spectra in MZ twins and unrelateds
Characterizing sleep spindles in 11,630 individuals from the National Sleep Research Resource
Purcell et al. (Nature Communications, 2017) [PubMed | PDF]
As a prelude to future molecular genetic studies, here we characterize the considerable variation, both between and within individuals, in sleep spindle activity.

Exome sequencing in schizophrenia
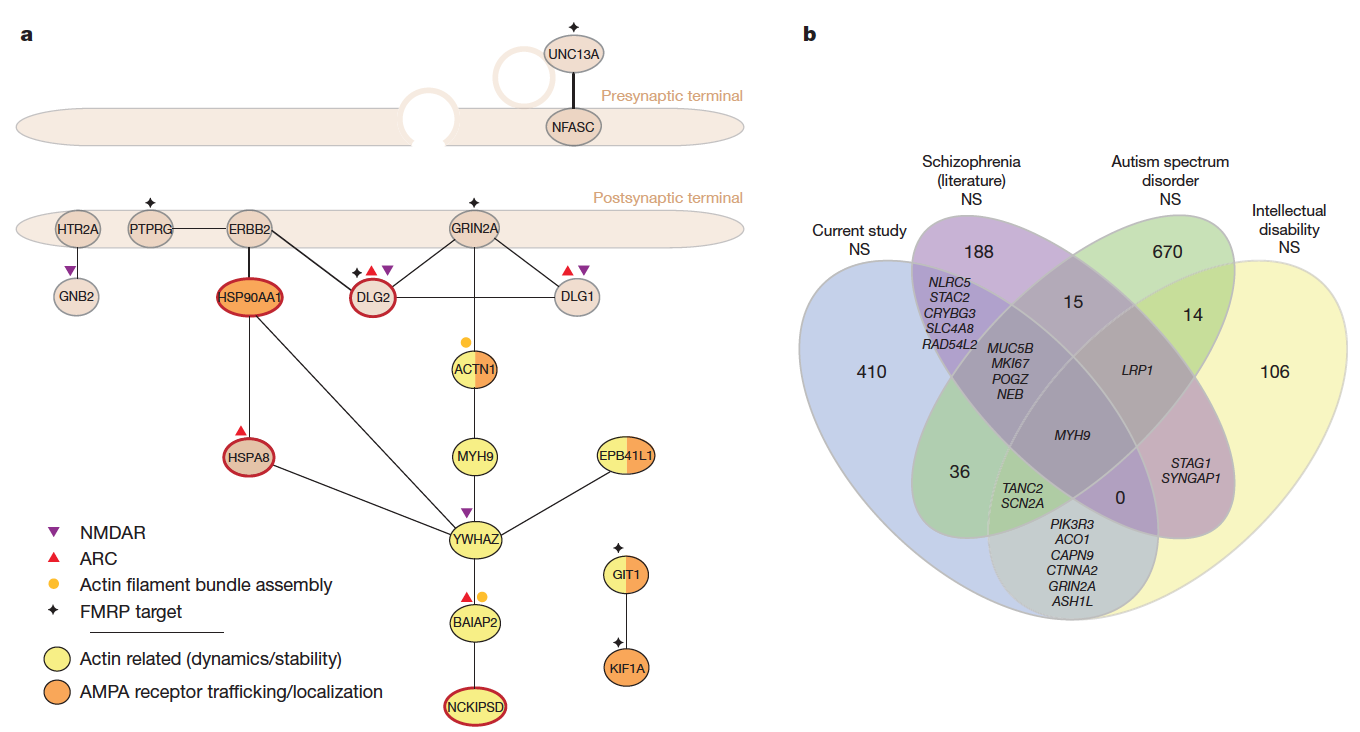
De novo mutations in schizophrenia

PLINK
PLINK: a tool set for whole-genome association and population-based linkage analyses
Purcell et al. (American Journal of Human Genetics, 2007) [PubMed | PDF]
This describes PLINK, a widely used software package for genome-wide association studies. We also introduced a novel approach to genome mapping based on identity-by-descent (IBD) information between very distantly related individuals.
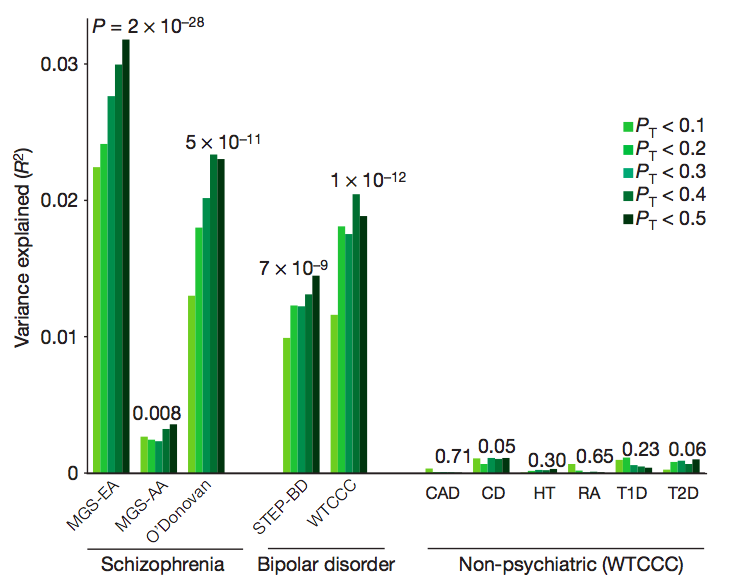
Polygenic risk profiles
Common polygenic variation contributes to risk of schizophrenia and bipolar disorder
International Schizophrenia Consortium (Nature, 2009) [PubMed | PDF]
One of the first large-scale genome-wide association studies of schizophrenia, here we made the observation that many common variants of very small effect appear to contribute to the heritability of schizophrenia. Most of these risk alleles are individually unlikely to be detectable, at least at currently feasible sample sizes.
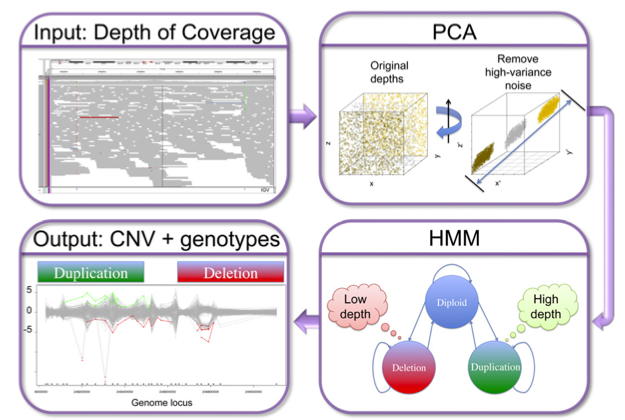
XHMM pipeline
Discovery and statistical genotyping of copy-number variation from whole-exome sequencing depth
Fromer et al. (American Journal of Human Genetics, 2012) [PubMed | PDF]
Reliably calling copy number variation from targeted sequencing data can be challenging. Here we present and evaluate a robust pipeline that overcomes many of the challenges that arise when trying to infer ploidy from sequencing read depth in exome sequencing studies.

Rare microdeletions
Rare chromosomal deletions and duplications increase risk of schizophrenia
International Schizophrenia Consortium (Nature, 2008) [PubMed | PDF]
Using genome-wide single nucleotide polymorphism arrays, we demonstrated an increased genome-wide burden of rare deletions and duplications (copy number variants, CNV) in patients with schizophrenia compared to controls. We also identified specific genomic loci at which CNVs pile up, namely 22q11, 15q13 and 1q21. [commentaries here and here]
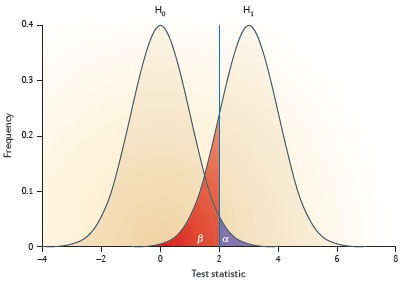
Statistical power and significance testing
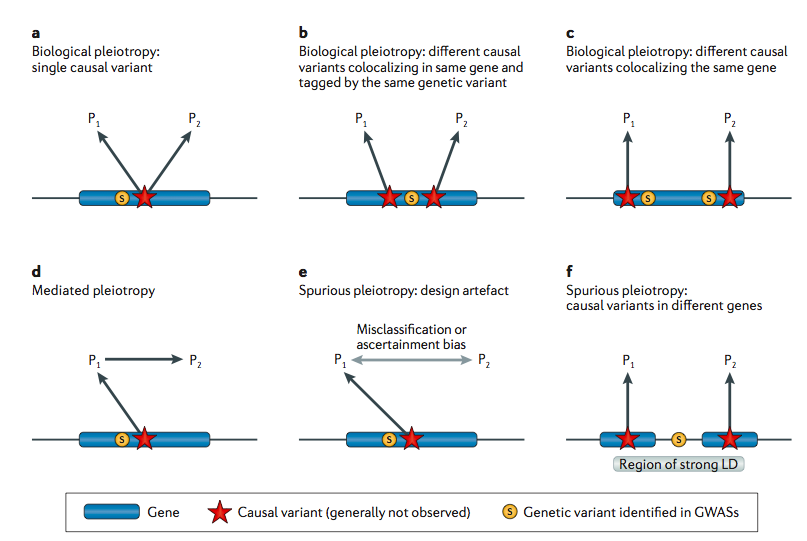
Models of pleiotropy
Pleiotropy in complex traits: challenges and strategies.
Solovieff et al. (Nature Reviews Genetics, 2013) [PubMed | PDF]
This Review considers approaches and interpretations of the often "one-to-many" nature of genotype-to-phenotype associations in complex disease. Nadia Solovieff was a postdoctoral researcher in the Purcell lab, jointly with Dr. Jordan Smoller's group.
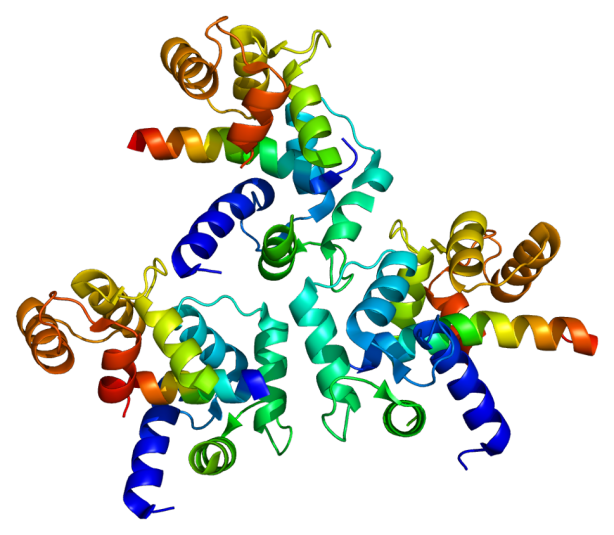
Calcium channel A1C sub-unit
Collaborative genome-wide association analysis supports a role for ANK3 and CACNA1C in bipolar disorder
Ferreira et al. (Nature Genetics, 2008) [PubMed | PDF]
This manuscript was the first meta-analysis of genome-wide single nucleotide polymorphism (SNP) data across studies of bipolar disorder. We implicated several genomic regions, including a role for calcium ion channel genes in this disease. [commentary]
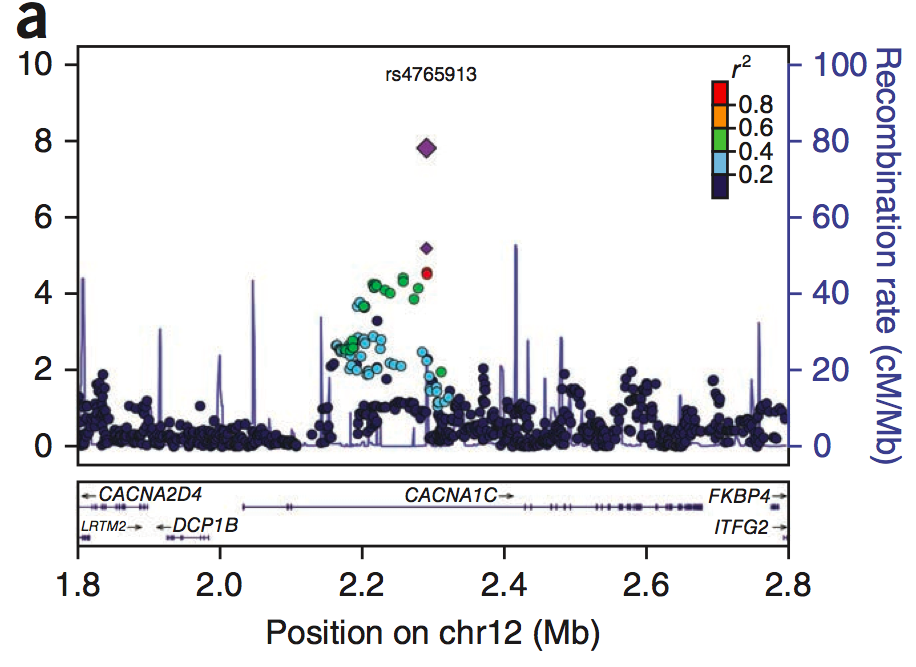
Large-scale genome-wide association analysis of bipolar disorder identifies a new susceptibility locus near ODZ4
Sklar et al. (Nature Genetics, 2012) [PubMed | PDF]
A later combined analysis of over 15,000 individuals, performed within the context of the PGC, provided further support for calcium channel and other genes.
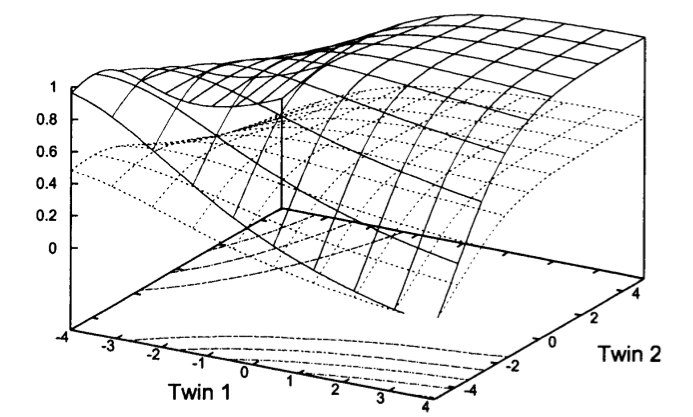
Variance components twin models
Variance components models for gene-environment interaction in twin analysis
Purcell (Twin Research, 2002) [PubMed | PDF]
A relatively old publication, included here to indicate some of our earlier work in behavioral genetics and twin studies. This manuscript provided a comprehensive methodological treatment for modeling how measured environmental variables can modify the relative impact of genetic influences for quantitative traits.
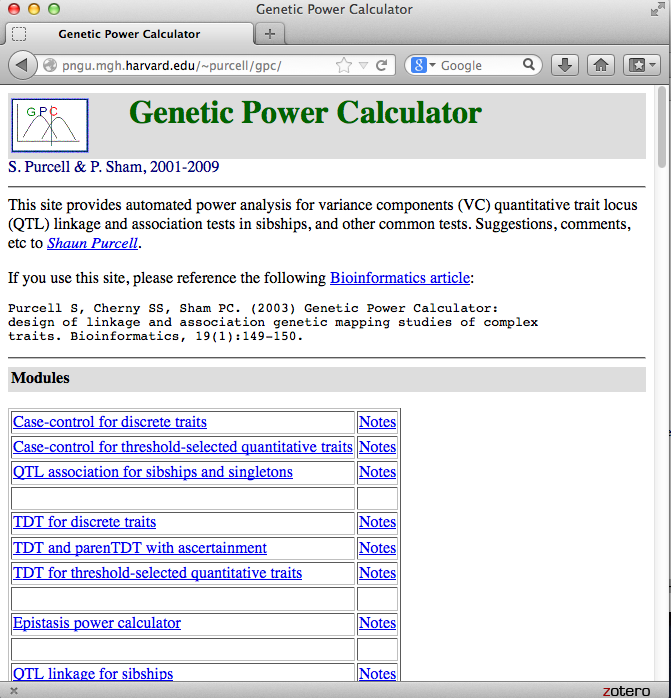
Genetic Power Calculator
Genetic Power Calculator: design of linkage and association genetic mapping studies of complex traits
Purcell et al. (Bioinformatics, 2003) [PubMed | PDF]
A small application note that describes a simple but widely used web tool for power calculation in the context of genetic association and linkage studies.