Epochs
Commands to define epochs for an EDF, and to attach annotations after loading an EDF
Epochs are fixed-length time windows into which Luna divides a recording for analysis — by default 30 seconds, matching AASM sleep staging convention. Most commands that operate on time-series data require epochs to be defined first, either explicitly with EPOCH or implicitly (Luna applies a 30-second default if none has been set). EPOCH also supports overlapping windows (e.g. 4-second epochs in 2-second steps) and annotation-based generic epochs. EPOCH-ANNOT loads an epoch-level annotation file in .eannot format, attaching stage or other labels to each epoch.
| Command | Description |
|---|---|
| Epoch definitions | Overview of epochs |
EPOCH |
Specify epochs |
EPOCH-ANNOT |
Load epoch-wise annotations from a .eannot-format file |
| Command table | Behavior of commands w.r.t. epochs & masks |
Epoch types
Epochs are time intervals that span the recording. Many Luna commands either explicitly or implicitly rely on epochs being defined. Epoching an EDF is often a first step that is required for many other Luna functions that work with epoched data.
In most instances, if epochs have not been specifically defined (by the EPOCH command), they will default to non-overlapping 30-second intervals, i.e. mirroring traditional (AASM) sleep staging.
By default, epochs are non-overlapping, of fixed duration (30 seconds) and contiguous with respect to clock-time (i.e. not spanning gaps in the recording). These epoch properties can be changed in various ways:
-
dur: epoch duration can be set to any duration in seconds, e.g. 5 seconds withdur=5 -
inc: the increment by default equalsdurimplying adjacent but non-overlapping epochs (e.g. 0-30 seconds, 30-60, 60-90, etc). Ifincis set to a different value, epochs can either be overlapping or (less likely) spaced. For example, withinc=15: 0-30, 15-45, 30-60, 45-75, etc).
TODO
- annot: generic__ or annotation-based_ epochs can be ... ___Fixed versus generic epochs: The most basic type of epoch is of fixed length (e.g. 30 seconds).
Epochs can be overlapping, e.g. shifting in 15 second increments
however, but by default, epochs are contiguous but not overlapping.
In contrast, generic epochs are those of potentially variable
duration, defined by existing annotations; this is
the most flexible form of epoch.
splice-gaps:
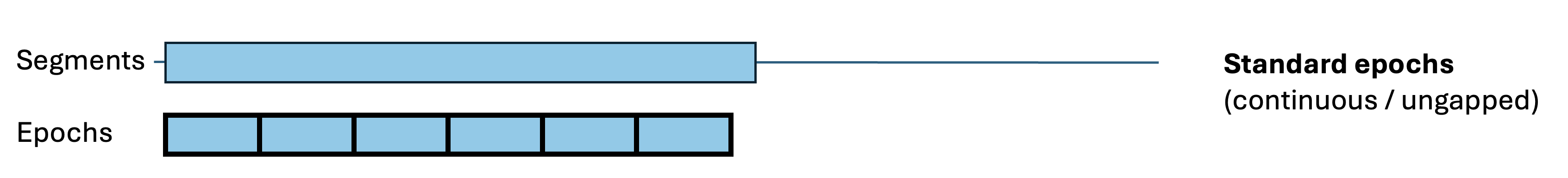
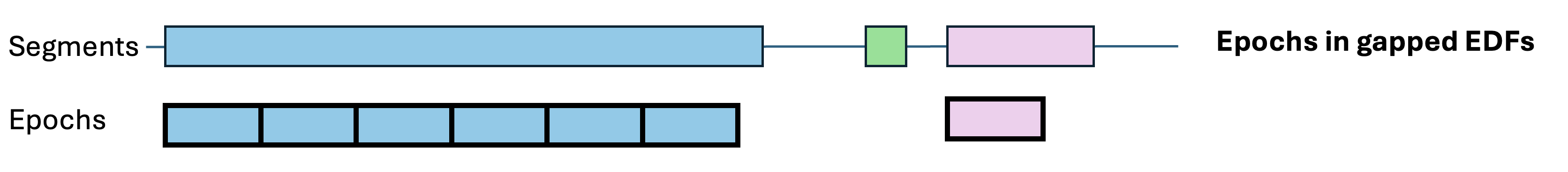
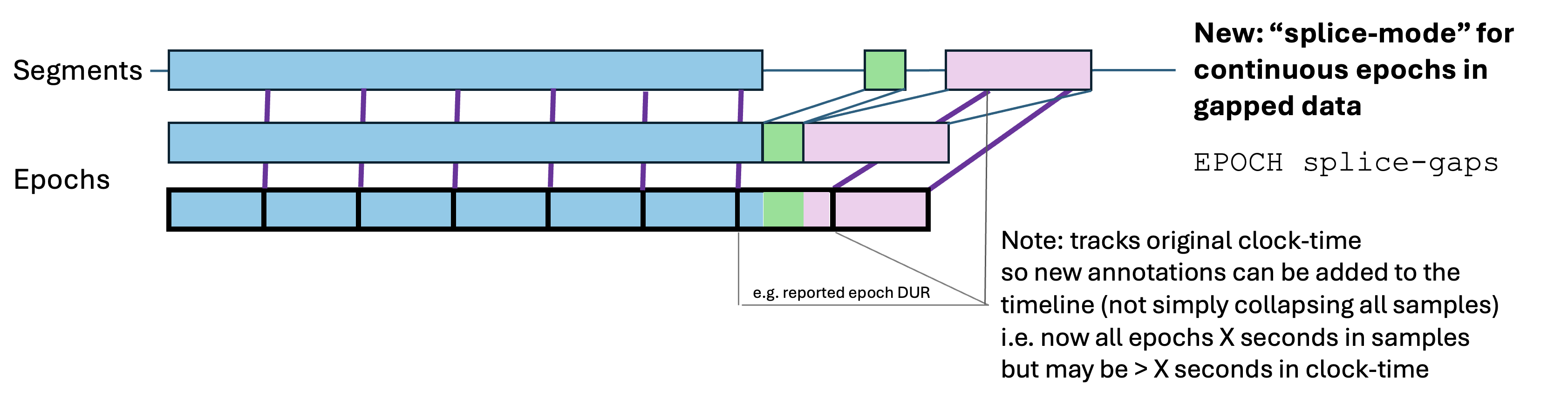
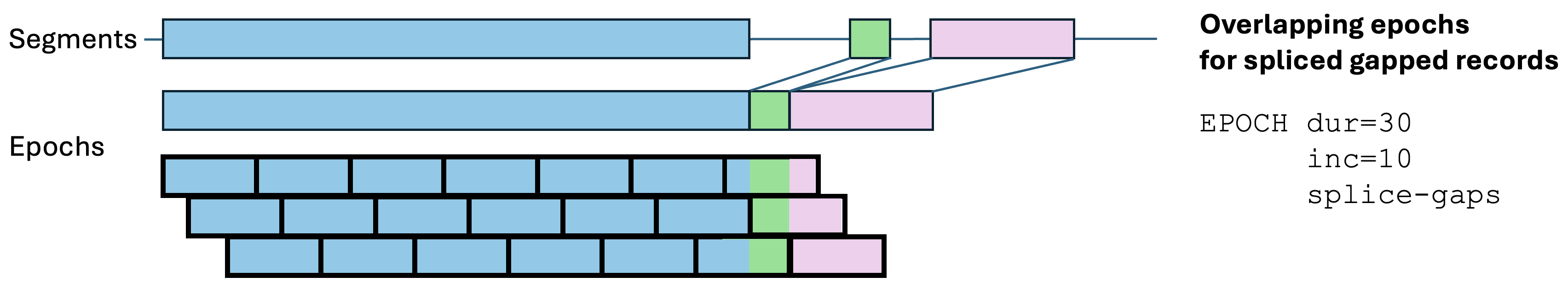
Some commands (in particular HYPNO) require standard,
non-overlapping epochs (otherwise, the resulting hypnogram metrics would
not be appropriately defined). Most but not all commands can work with generic
epochs, as detailed below. Luna will give an error message if trying
to use generic epochs with a command that does not accept them.
EPOCH
Divides the time series into epochs
This command can define either standard or generic epochs.
Parameters
Without any parameters set, the default is 30-seconds, with no overlap
(i.e. 30-second increments, equivalent to len=30 or epoch=30). In
general, if the increment is not specified, it defaults to the epoch
length (i.e. no overlap between epochs).
By default, epochs start at 0 seconds (start of EDF).
| Parameter | Example | Description |
|---|---|---|
len |
30 | Epoch length (seconds), defaults to 30 |
inc |
30 | Epoch increment (seconds), defaults to len (i.e. no overlap) |
epoch |
30,15 | Epoch length{,increment} (seconds), defaults to 30,30 |
require |
10 | Stop processing that EDF if there are not at least N epochs |
align |
N1,N2,N3,R,W,? |
Align epoch starts with the first of these annotations |
offset |
10 | Explicitly set start time for first epoch (seconds) |
clear |
Reset any defined epochs |
Note: if align is not given an explicit argument, it defaults to N1,N2,N3,R,W,?,L,U,M - i.e.
a list of common stage annotations.
Output options
| Parameter | Example | Description |
|---|---|---|
min |
Minimal output: writes the number of epochs to standard output | |
verbose |
Output epoch-level information when setting epochs | |
table |
As verbose except do not change the current epochs, only output the table |
Generic epochs
| Parameter | Example | Description |
|---|---|---|
annot |
spindle |
Set generic epochs based on existing annotations |
w |
5 |
Expanded window (seconds) both before and after each annotation |
w-before |
5 |
Extend annotation w-before seconds before the start point |
w-after |
5 |
Extend annotation w-after seconds after the stop point |
start |
Reduce annotations to their start (designed to work with w ) |
|
midpoint |
Reduce annotations to their midpoint (designed to work with w ) |
|
stop |
Reduce annotations to their stop (designed to work with w ) |
Outputs
Basic summary information (strata: none)
| Variable | Description |
|---|---|
DUR |
Epoch duration (as per the input) |
INC |
Epoch increment (as per the input) |
NE |
Number of epochs in the EDF, given DUR and INC |
GENERIC |
Flag to indicate whether epochs are generic (0/1) |
OFFSET |
Any offset (from EDF start) |
Per-epoch interval information (option: verbose, strata: E)
| Variable | Description |
|---|---|
E1 |
Current epoch number (which may differ from E if the EDF has been restructured) |
DUR |
Epoch duration (secs) |
LABEL |
Epoch label |
HMS |
Clock-time for epoch start (hh:mm:ss) |
INTERVAL |
String label of epoch interval (seconds) |
MID |
Midpoint of epoch (seconds elapsed from EDF start) |
START |
Start of epoch (seconds elapsed from EDF start) |
STOP |
Stop of epoch (seconds elapsed from EDF start) |
TP |
Epoch interval in time-points (1TP = 1e-9 sec) |
Example
Fixed epochs
To set epochs of 10-seconds, each with a 5-second overlap (output showing the first six epochs):
luna my.edf -o out.db -s ' EPOCH len=10 inc=5 verbose '
To get the number of epochs set:
destrat out.db +EPOCH
ID DUR GENERIC INC NE OFFSET
nsrr01 10 0 5 8183 0
destrat out.db +EPOCH -r E
ID E DUR E1 HMS INTERVAL LABEL MID START STOP TP
nsrr01 1 10 1 21:58:17 0.00->10.00 E1 5 0 10 0->10000000000
nsrr01 2 10 2 21:58:22 5.00->15.00 E2 10 5 15 5000000000->15000000000
nsrr01 3 10 3 21:58:27 10.00->20.00 E3 15 10 20 10000000000->20000000000
nsrr01 4 10 4 21:58:32 15.00->25.00 E4 20 15 25 15000000000->25000000000
nsrr01 5 10 5 21:58:37 20.00->30.00 E5 25 20 30 20000000000->30000000000
nsrr01 6 10 6 21:58:42 25.00->35.00 E6 30 25 35 25000000000->35000000000
... (etc) ...
If we instead first applied a MASK (which implicitly run EPOCH
if the data aren't already epoched), RESTRUCTUREed the data,
and then requested information only (table) from EPOCH:
MASK epoch=10-14 & RE & EPOCH table
E and E1):
ID E DUR E1 HMS INTERVAL LABEL MID START STOP TP
nsrr01 10 30 1 22:02:47 270.00->300.00 E10 285 270 300 270000000000->300000000000
nsrr01 11 30 2 22:03:17 300.00->330.00 E11 315 300 330 300000000000->330000000000
nsrr01 12 30 3 22:03:47 330.00->360.00 E12 345 330 360 330000000000->360000000000
nsrr01 13 30 4 22:04:17 360.00->390.00 E13 375 360 390 360000000000->390000000000
nsrr01 14 30 5 22:04:47 390.00->420.00 E14 405 390 420 390000000000->420000000000
Info
The above command implicitly ran an EPOCH command prior to the MASK; the
above output will be from the second EPOCH. Here, using
TAGs can keep track of the various outputs: e.g.
TAG R/1 & EPOCH & MASK epoch=10-14 & RE & TAG R/2 & EPOCH table
EPOCH:
destrat out.db +EPOCH -r R
ID R DUR GENERIC INC NE OFFSET
nsrr01 1 30 0 30 1364 0
nsrr01 2 30 0 30 5 0
NE.
Generic epochs
Generic epochs are those based on existing annotations. One or more
annotation labels follow the annot option of EPOCH: e.g.
EPOCH annot=Obstructive_Apnea,Hypopnea
will define epochs based on apnea or hypopnea events. Note that
generic epochs can be a variable size, and can overlap. Running the above (with verbose)
produces the following outputs:
destrat out.db +EPOCH
ID GENERIC NE
nsrr02 1 106
GENERIC is now set to 1, but there is no information on epoch duration - i.e. because it may vary between epochs. That information
is available in the epoch-stratified output (requested by verbose):
destrat out.db +EPOCH -r E
ID E DUR E1 HMS INTERVAL LABEL MID START STOP TP
nsrr02 1 10.8 1 22:08:49 3043.70->3054.50 Hypopnea 3049.1 3043.7 3054.5 3043700000000->3054500000000
nsrr02 2 10.5 2 22:16:53 3527.30->3537.80 Hypopnea 3532.55 3527.3 3537.8 3527300000000->3537800000000
nsrr02 3 13 3 22:53:44 5738.40->5751.40 Hypopnea 5744.9 5738.4 5751.4 5738400000000->5751400000000
nsrr02 4 10.5 4 22:55:43 5857.90->5868.40 Hypopnea 5863.15 5857.9 5868.4 5857900000000->5868400000000
nsrr02 5 16.1 5 22:56:01 5875.30->5891.40 Hypopnea 5883.35 5875.3 5891.4 5875300000000->5891400000000
nsrr02 6 15.9 6 22:59:33 6087.20->6103.10 Hypopnea 6095.15 6087.2 6103.1 6087200000000->6103100000000
... (etc) ...
DUR is the duration of each epoch (which here matches the duration of each hypopnea annotation). The LABEL is
the annotation name (Hypopnea or Obstructive_Apnea).
Re-running with the altered command:
EPOCH annot=Obstructive_Apnea,Hypopnea verbose start w-after=5
ID E DUR E1 HMS INTERVAL LABEL MID START STOP TP
nsrr02 1 5 1 22:08:49 3043.70->3048.70 Hypopnea 3046.2 3043.7 3048.7 3043700000000->3048700000000
nsrr02 2 5 2 22:16:53 3527.30->3532.30 Hypopnea 3529.8 3527.3 3532.3 3527300000000->3532300000000
nsrr02 3 5 3 22:53:44 5738.40->5743.40 Hypopnea 5740.9 5738.4 5743.4 5738400000000->5743400000000
nsrr02 4 5 4 22:55:43 5857.90->5862.90 Hypopnea 5860.4 5857.9 5862.9 5857900000000->5862900000000
nsrr02 5 5 5 22:56:01 5875.30->5880.30 Hypopnea 5877.8 5875.3 5880.3 5875300000000->5880300000000
nsrr02 6 5 6 22:59:33 6087.20->6092.20 Hypopnea 6089.7 6087.2 6092.2 6087200000000->6092200000000
... (etc) ...
DUR is fixed at 5 seconds for each epoch; note how the start times are the same as above (e.g. 3043.70 seconds for the first epoch)
but now the end is altered, reflecting the effect of start w=5.
Reducing/expanding annotations to epochs
If requested, note
that either start, midpoint or stop are applied first
(i.e. shrinking the annotation to a single time-point), followed
by either w, w-before or w-after (i.e. expanding the
interval either before, after or around (centered on) that
time-point. Note that it is also possible to apply the w*
expansion options by first reducing the annotations by start,
midpoint or stop. This expands the interval by the requested number
of seconds, at either the start (w-before), the end (w-after) or at both
start and end (w).
EPOCH-ANNOT
Attach epoch-level annotations from a file, to an epoched EDF
This command reads a plain-text file from disk, expecting the
.eannot format. It is possible to
apply multiple sets of annotations to epochs with multiple
EPOCH-ANNOT commands.
The behavior of this command is similar but not identical to what would
happen if the .eannot file were directly specified in the
sample-list (i.e. as any other XML,
.annot annotation file). As this command can be performed
after the EDF has been loaded and manipulated (e.g. via
RESTRUCTURE), the number of epochs
(i.e. rows in the file) should match exactly the number of epochs
given the current state of the in-memory EDF (i.e. after any
restructuring, and with the current epoch definitions).
That is, if EPOCH generates 1022 epochs for a given EDF, then
the annotation file must be exactly 1022 epochs. Luna will give an
error if the EPOCH command has not been performed prior to
EPOCH-ANNOT.
In contrast, when attaching an .eannot file via the sample-list,
Luna assumes the .eannot file corresponds to the entire on-disk
EDF (typically with the default 30-second epochs). (When attaching
initially from a sample list (not EPOCH-ANNOT) the tolerance
for exact matching can be altered with the
special variables
no-epoch-check and epoch-check.)
An optional recode parameter can be used to translate annotations on-the-fly.
Make sure to use plain-text files
Annotation files should be plain-text ASCII files with no special encodings, not RTF (rich text format), etc. Please see this FAQ.
Parameters
| Parameter | Example | Description |
|---|---|---|
file |
file=annots/id1.epochs |
File path/name to read annotations from (required) |
recode |
recode=ST1=N1,ST2=N2 |
Optional, comma-delimited list of recodings (from=to) |
Outputs
This command does not produce any output per se, other than noting the number of epoch/annotations read in the log/console, e.g.
CMD #2: EPOCH-ANNOT
mapping 2 distinct epoch-annotations (1022 in total) from annots/test.eannot
Example
Here we attach epoch-annotations to an epoched EDF and use them to
mask out certain epochs. For example, here we have a
dummy annotation file, that contains an annotation of either T or
A for each epoch. This particular EDF has 1022 30-second epochs,
and so the file epoch1.eannot is the same length:
$ sort epoch1.eannot | sort | uniq -c
102 A
920 T
The file epoch1.eannot is a simple text-file, with either an
A or a T on each line, and nothing else.
Note
Annotations are not restricted to A or T or single
characters: they can be any text string, but should not contain
spaces or special characters. A common use may be if sleep
staging has been performed, so each row indicates N1, N2,
N3, REM or Wake, etc.
To attach this file, and use it to select only A epochs:
luna my.edf -s ' EPOCH & EPOCH-ANNOT file=epoch1.eannot & MASK ifnot=A & RE '
..................................................................
CMD #1: EPOCH
set epochs, length 30 (step 30), 1022 epochs
..................................................................
CMD #2: EPOCH-ANNOT
mapping 2 distinct epoch-annotations (1022 in total) from epoch1.eannot
..................................................................
CMD #3: MASK
set masking mode to 'mask' (default)
based on A 102 epochs match; 920 newly masked, 0 unmasked, 102 unchanged
total of 102 of 1022 retained
..................................................................
CMD #4: RESTRUCTURE
restructuring as an EDF+ : keeping 3060 records of 30664, resetting mask
retaining 102 epochs
Having selected epochs based on these external annotations,
one might subsequently apply other analysis commands, or write out a new EDF (see
WRITE), for example.
Working with multiple EDFs
When using sample-lists to
work with multiple EDFs, it is often convenient to use the ^
special character in a script, which is replaced with the ID (from
the first column of the sample-list, not the EDF header) for that
individual. This enables the same script to point to different
annotations files for different individuals (assuming
that those annotation files are named based on the IDs).
Here is an example of using a script (instead of the -s option) to
achieve the same result as above, but here using a sample list and
renaming the annotation file to, for example,
annots/id001.epoch.eannot. We might specify the sample list s.lst
as follows, to assign the ID id001 to the EDF:
id001 my.edf
The Luna script (extract.txt) below
also illustrates using a variable
to specify more flexibly the required annotation,
and writing a new EDF (with the WRITE command):
% Epoch the data (default 30 sec)
EPOCH
% Attach the matching file in the annots/ folder for this individual
% i.e. requires that files are named
EPOCH-ANNOT file=annots/^.eannot
% Set this mask, where the value of ${x} will be specified
% on the command line
MASK ifnot=${x}
% Apply the mask, actually restructuring the internal EDF
% (Note, RESTRUCTURE can be abbreviated as RE)
RESTRUCTURE
% Write a new EDF back to disk
WRITE edf-dir=edfs/ % write new EDFs in this directory/folder
edf-tag=extract-${x} % add this tag to the filename for each EDF
sample-list=s-${x}.lst % make a new sample-list to point to the new EDFs
Note how the ^ character is used to specify the individual/EDF
ID. We can then run Luna, setting the x variable to A:
luna s.lst x=A < extract.txt
For individual id001, for example, Luna searches for
annots/id001.eannot and then masks epochs that do not match the
value specified by the variable ${x} (here set to A).
After RESTRUCTURE-ing, it WRITEs an EDF to edfs/my-extract-A.edf
and creates a matching sample list s-A.lst. As a sanity check,
running DESC on the new project:
luna s-A.lst -s DESC
EDF file : edfs/my-extract-A.edf
ID : id001
Duration : 00:51:00
Number of signals : 6
Signals : EOG-L[256] EOG-R[256] EMG[256] EEG1[256] EEG2[256] EEG3[256]
Command & epoch types
Not all commands can be combined with generic epochs. Here we list the current constraints of particular commands with respect to epoch definitions. These fall into the following classes:
-
commands that ignore epoch definitions
-
commands that work with any epoch definition
-
commands that cannot use generic epoch definitions
Generally, Luna will tell you if there is a problem, but the detailed tables below are included for reference.
Commands that ignore epoch definitions
The following primarily do not touch signal-level data and are therefore not impacted by the presence or absence of any type of epoching:
ALIASES ALIGN ANNOTS ANON CANONICAL CLOCS CONTAINS
CPT CWT-DESIGN DESC EDF ENFORCE-SR FILTER-DESIGN HEADERS
MAKE-ANNOTS MEANS OVERLAP RECORD-SIZE REMAP RENAME
SET-HEADERS SET-TIMESTAMPS SET-VAR SIGGEN SIGNALS
SIMUL SL SUMMARY TAG TIME-TRACK TYPES VARS WRITE-ANNOTS
The following pull the entire signal and treat it as a continuous time-series - i.e. they effectively ignore any epoch definitions. Typically, commands that manipulate/generate an EDF signal do not work with epochs, as they must replace the whole signal in the EDF.
1FNORM A2S ACF ADJUST ALTER COPY CWT DUMP-RECORDS
EMD FFT FILTER FLIP FREEZE HILBERT ICA MINMAX
mV OTSU RECS RECTIFY REFERENCE RESAMPLE
REVERSE S2A SEGMENTS SPANNING
THAW TLOCK TRANS TV uV WRITE Z-PEAKS
Commands that work with any epoch definitions
These commands will not consider signal data outside of unmasked epochs.
Of these, the following allow generic (variable-sized) epochs:
| Command | Notes |
|---|---|
CHEP |
Variable epoch size allowed, but not directly accounted for in analysis |
CHEP-MASK |
(makes more sense w/ equal sized : epochs weighted equally) |
DUMP-MASK |
|
DUPES |
|
EPOCH |
|
EPOCH-ANNOT |
If no epochs defined, adds as fixed default (30s) |
GP |
|
HEAD |
Select which epoch w/ "epoch" arg; always dump from epoch |
INTERPOLATE |
|
MASK |
|
MATRIX |
Only outputs epoched data |
MSE |
|
MTM |
|
PSC |
Only works on stats, which may be epoch level but size etc is arbitrary; they will be equally weighted in analyses however |
PSD |
|
RESTRUCTURE / RE |
|
SEDF |
(but drops epoch timing information in output .sedf) |
SIGSTATS |
Outputs simple, unweighted epoch means |
TABULATE |
|
XCORR |
|
| ``` |
Allow whole-recording, or epoch-level analysis, and allow variable-sized epochs:
| Command | Notes |
|---|---|
CORREL |
(w/ epoch or ch-epoch) |
EVAL |
interval for whole-recording |
EXE |
(w/ epoch) |
LZW |
(w/ epoch) |
MI |
(w/ epoch) |
MS |
(w/ epoch) |
PSI |
(w/ epoch) |
SO |
Extra output only |
SPINDLES |
Assumes fixed epoch length for spindle/SO shuffling within-epoch (gives warning/error) : otherwise okay (for basic spindle counting per epoch) |
STATS |
(w/ epoch) |
Commands that only work with uniform-sized epochs
Epoch-level commands that do not allow variable-sized epochs:
| Command | Notes |
|---|---|
ARTIFACTS |
|
EVAL-STAGES |
|
HYPNO |
|
PLACE |
|
POPS |
|
REBASE |
|
SOAP |
|
STAGE |
These commands can be run in either whole-signal mode, but if run in epoch-level mode, they do not allow variable-sized epochs:
| Command | Notes |
|---|---|
CC |
w/ epoch |
COH |
w/ epoch |
IRASA |
Ignores any offset; assumes fixed epoch len, no overlap + zero offset |
LINE-DENOISE |
Assumes epochs are contiguous and no offset (and spans whole record, or last few points uncorrected) |
MOVING-AVERAGE |
w/ epoch, assumes epochs are contiguous and no offset (and spans whole record, or last few points uncorrected) |
PEAKS |
w/ epoch, assumes contiguous epochs |
ROBUST-NORM |
w/ epoch, assumes epochs are contiguous and no offset (and spans whole record, or last few points uncorrected) |
SUPPRESS-ECG |
Command currently forces 30s epochs after pulling whole signal |
ZC |
w/ epoch, assumes epochs are contiguous and no offset (and spans whole record, or last few points uncorrected) |