Manipulations
Commands to alter basic properties of the EDF and the signals therein
This page covers a broad set of commands for modifying signals and EDF properties in memory. Channel management commands (SIGNALS, RENAME, COPY) control which channels are present and how they are labelled. Unit and amplitude commands (uV, mV, REFERENCE, FLIP, ZC, ROBUST-NORM, ROLLING-NORM, CLIP, SCALE, LOG, RECTIFY) adjust signal values. Timing and structural commands (RESAMPLE, RECORD-SIZE, EDF-MINUS) modify sample rates and EDF record layout. Header and metadata commands (ANON, SET-HEADERS, SET-VAR, SET-TIMESTAMPS) update EDF header fields and Luna variables. These tools are typically used to clean and standardize EDFs before running analytical commands.
| Command | Description |
|---|---|
SIGNALS |
Retain/remove specific EDF channels |
RENAME |
Rename channels |
COPY |
Duplicate one or more EDF channels |
RESAMPLE |
Resample signal(s) |
ENFORCE-SR |
Require a particular sample rate |
REFERENCE |
Re-reference signals |
DEREFERENCE |
De-reference signals |
MINMAX |
Set digital/physical min/max across channels |
uV |
Rescale units to uV |
mV |
Rescale units to mV |
FLIP |
Flip polarity of signal |
ZC |
Mean-center signal |
ROBUST-NORM |
Robust normalization |
CLIP |
Clip a signal using absolute or relative thresholds |
COMBINE |
Combine two or more channels into a new channel (e.g. sum/mean) |
SCALE |
Rescale a channel (min/max scaling) |
LOG |
Apply a log transform to one or more signals |
ROLLING-NORM |
Rolling-window normalization |
SHIFT |
Shift a signal |
SCRAMBLE |
Scramble a signal |
TIME-TRACK |
Add a time-track to an EDF |
RECORD-SIZE |
Change EDF record size |
EDF-MINUS |
Realign EDF records, annotations and epochs |
INSERT |
Align and splice signals from a secondary EDF, with clock-drift estimation |
ANON |
Strip ID information from EDF header |
SET-HEADERS |
Directly specify certain EDF headers |
SET-VAR |
Directly specify Luna variables |
SET-TIMESTAMPS |
Directly specify EDF record timestamps |
RECTIFY |
Rectify a signal |
REVERSE |
Reverse a signal |
MOVING-AVERAGE |
Moving average (or median) of a signal |
SIGNALS
The command requires one of two options: either keep or drop.
Each expects a comma-delimited list of channel names (or
aliases), which are either retained or
removed from the in-memory dataset.
Methods
Specified channels are selectively retained or removed from the in-memory recording, reducing the dataset to only those signals required for subsequent analysis steps.
Parameters
| Option | Example | Description |
|---|---|---|
drop |
drop=EMG,ECG |
Drop channels EMG and ECG |
keep |
keep=C3,C4 |
Drop all channels except C3 and C4 |
Outputs
Other than modifying the in-memory representation of the EDF, there is no further output (except some notes written to the log).
Example
For an EDF with 6 signals, including EMG, EOG-L and EOG-R, this command would drop these three signals:
luna s.lst -s 'SIGNALS drop=EMG,EOG-L,EOG-R & DESC'
DESC:
Number of signals : 3
Signals : EEG1[256] EEG2[256] EEG3[256]
keep option with the same arguments:
luna s.lst -s 'SIGNALS keep=EMG,EOG-L,EOG-R & DESC'
Number of signals : 3
Signals : EOG-L[256] EOG-R[256] EMG[256]
RENAME
Renames channels
This command can rename channels within the context of evaluating a Luna script. In this way, it differs from using signal aliases (which can only be specified when first initiating Luna, and will be fixed for all individuals), as this command can use variables (which may be individual-specific).
It is also possible to supply a file of new labels for multiple signals instead.
Methods
Channel labels are reassigned within the in-memory representation of the recording, allowing recording-specific naming conventions to be mapped to a standardized nomenclature. Unlike static pre-processing aliases, relabelling at this stage allows individual-specific label assignments to be applied dynamically within a processing script.
Parameters
| Parameter | Example | Description |
|---|---|---|
sig |
C3,C4 |
List of channels to duplicate |
new |
C3_LM,C4_LM |
List of new labels (same size as sig) |
file |
f.txt |
File of old/new labels |
Note that you cannot use an existing channel label as a new label.
If specifying a file, such files should be tab-delimited, containing exactly
two fields per row (original label as per sig)
and the new value as per new).
Output
No formal output, other than changing the labels of channels in the internal EDF
Example
In its simplest form, if we have a channel named THOR_RES, for
example, we can rename it to some other label -- here just using XX --
using RENAME, and then use that new label in other commands:
luna s.lst -s ' RENAME sig=THOR_RES new=XX & STATS sig=XX '
Note that this also adds an alias internally, so that THOR_RES can
still be used as a label (e.g. with the sig option of a command),
but the output will be labelled with the primary term XX.
As noted above, this provides similar functionality as using a signal alias:
luna s.lst "alias=XX|THOR_RES" -s ' STATS sig=XX '
The primary difference is that RENAME accepts (individual-specific)
variables as arguments, i.e. which can allow different individuals to
have different assignments (with sig and/or new). For example, in
this toy example, we change THOR_RES and ABDO_RES (all present in
the three individuals in the tutorial dataset) to
different labels. If we have a tab-delimited file that defines these
variables for each individual:
cat ch.txt
ID CHS
nsrr01 XX,YY
nsrr02 AA,BB
nsrr03 CC,DD
Now, the command
luna s.lst vars=ch.txt \
-s ' DESC & RENAME sig=THOR_RES,ABDO_RES new=${CHS} & DESC '
will use RENAME to swap those two channels to the other specified
values: e.g. showing extracts from the DESC output for the three
individuals: in all cases, the first DESC command gives the same:
Signals : SaO2[1] PR[1] EEG_sec_[125] ECG[250] EMG[125] EOG_L_[50]
EOG_R_[50] EEG[125] AIRFLOW[10] THOR_RES[10] ABDO_RES[10] POSITION[1]
LIGHT[1] OX_STAT[1]
DESC varies between the three individuals as expected:
Signals : SaO2[1] PR[1] EEG_sec_[125] ECG[250] EMG[125] EOG_L_[50]
EOG_R_[50] EEG[125] AIRFLOW[10] XX[10] YY[10] POSITION[1]
LIGHT[1] OX_STAT[1]
Signals : SaO2[1] PR[1] EEG_sec_[125] ECG[250] EMG[125] EOG_L_[50]
EOG_R_[50] EEG[125] AIRFLOW[10] AA[10] BB[10] POSITION[1]
LIGHT[1] OX_STAT[1]
Signal : SaO2[1] PR[1] EEG_sec_[125] ECG[250] EMG[125] EOG_L_[50]
EOG_R_[50] EEG[125] AIRFLOW[10] CC[10] DD[10] POSITION[1]
LIGHT[1] OX_STAT[1]
Another difference is that using signal aliases allows a many-to-one
mapping, whereas RENAME requires a one-to-one mapping of
labels. That is, "alias=XX|AA|BB|CC" will map either AA, BB or
CC to XX (i.e. where an individual EDF may have none, one or
multiple of these labels). In contrast, RENAME must use the single,
primary label in the sig option. However, there is nothing stopping
combined use of signal aliases and the RENAME command, i.e. to achieve a many-to-many
mapping of labels: e.g.
luna s.lst vars=ch.txt "alias=XX|AA|BB|CC" \
-s ' RENAME sig=XX new=${CHS} & WRITE edf-dir=edfs/ '
This effectively uses XX as an intermediate (mapped from either AA, BB or CC) and will then write to
the new EDF a label as defined in ch.txt. Of course, an alternative would be to also supply individual-specific
labels for both sig and new in the vars.txt file, e.g. if it had two columns defining OLD and NEW variables/columns:
luna s.lst vars=ch.txt \
-s ' RENAME sig=${OLD} new=${NEW} & WRITE edf-dir=edfs/ '
ROLLING-NORM
Normalize a signal using a rolling local window
ROLLING-NORM applies a rolling-window normalization to one or more signals
across the whole retained trace. The command uses a local window around each
sample to compute a mean and standard deviation, then rescales the central
sample to a local z-score. This can be useful when a signal has slow drift or
time-varying scale that is not well handled by a single whole-trace
normalization.
Methods
For each requested signal, Luna reads the whole retained trace and computes a
locally normalized version using a symmetric rolling window of width w
seconds. Internally, the window size is converted to samples using the signal's
sample rate and forced to an odd number. At each sample, the central value is
standardized relative to the samples in that local window, producing a
time-varying z-score. The normalized trace replaces the original in-memory
signal.
Parameters
| Parameter | Example | Description |
|---|---|---|
sig |
sig=C3,C4 |
Signals to normalize |
w |
w=30 |
Rolling window width in seconds; must be at least 1 second |
Output
No formal tabular output. The command updates the in-memory signal values.
Example
luna s.lst -s 'ROLLING-NORM sig=EEG w=30'
COPY
Duplicates one or more EDF channels
Because some Luna commands modify a channel
(e.g. FILTER), it can be desirable to first
make a copy of the original channel. New channels are written out
with the WRITE command.
Although multiple signals can be duplicated at the same time (i.e. will all be given the same tag), only data channels (i.e. not EDF Annotation channels in EDF+) are duplicated.
Methods
One or more channels are duplicated within the in-memory recording, preserving the original signal while allowing subsequent processing steps (such as filtering or normalization) to be applied to the copy without altering the source channel.
Parameters
| Parameter | Example | Description |
|---|---|---|
sig |
sig=C3,C4 |
List of channels to duplicate |
tag |
tag=DELTA |
A required option, this is added to make the new channel name, e.g. C3 becomes C3_DELTA |
Output
One or more new channels are created in the in-memory representation of the EDF. Aside from a note in the log, there is no formal (destrat-based) output for this command.
Example
To extract one channel (EEG) from an original EDF, and then duplicate it:
luna s.lst 2 sig=EEG -s 'DESC & COPY sig=EEG tag=V2 & DESC'
As expected, the first DESC output shows a single channel:
EDF filename : edfs/learn-nsrr02.edf
ID : nsrr02
Clock time : 21:18:06 - 07:15:36
Duration : 09:57:30
# signals : 1
Signals : EEG[125]
After the COPY command has been executed, there are now two channels: EEG and EEG_V2:
EDF filename : edfs/learn-nsrr02.edf
ID : nsrr02
Clock time : 21:18:06 - 07:15:36
Duration : 09:57:30
# signals : 2
Signals : EEG[125] EEG_V2[125]
RESAMPLE
Changes the sampling rate of a signal
Uses functions from libsamplerate to upsample or downsample signals.
Within a maximum upsampling/downsampling factor of 256, there are no
constraints on the new sample rate (i.e. the ratio of old and new
sample rates need not be a rational number).
The downsample and upsample-if options can be used to control whether a channel is resampled:
-
downsamplemeans that channels will only ever be downsampled, otherwise left as is -
upsample-if=50means that channels will only be resampled (up or down) if the rate is above, e.g. 50 Hz. This option can avoid "garbage channels" (e.g. a channel labelled as an EEG but with a sample rate of 10 Hz, for example) being included.
Methods
Signals are resampled to a common target rate using a high-quality sinc-interpolation algorithm (via the Secret Rabbit Code / libsamplerate library). The approach supports arbitrary rational and irrational resampling ratios, applying anti-aliasing lowpass filtering prior to downsampling and sinc-based interpolation for upsampling, thereby minimising aliasing and interpolation artefacts. Channels can be selectively downsampled only (leaving channels already at or below the target rate unmodified) or restricted to those above a minimum rate threshold to prevent unintended resampling of low-rate auxiliary channels.
Parameters
| Parameter | Example | Description |
|---|---|---|
sig |
sig=C3,C4 |
Signal list |
sr |
sr=128 |
New sampling rate (Hz) |
downsample |
Only channels with rates above sr will be downsampled |
|
upsample-if |
upsample-if=100 |
Only resample if the original sample rate is greater than this value |
Output
No output other than a message to the log (and altering the in-memory signal).
Example
To create a new EDF with the EEG channel resampled to 100 Hz:
luna s.lst -s 'RESAMPLE sig=EEG sr=100 & WRITE edf-tag=resample edf-dir=edfs/ sample-list=s2.lst'
ENFORCE-SR
Drop signals that do not conform to a given EDF record size, or min/max sample rate
To enable clean EDF record-size conversion, this command first drops
any signals which would not be able to be represented by an N-second
record size. For example, a signal with a sample rate of 125 Hz cannot
be represented in the EDF block/record structure if each block is only
0.5 seconds. Such signals will be dropped (based on dur) from the
internal EDF representation. This command can be used prior to a
RECORD-SIZE command.
Further, this command can optionally drop signals with a sample rate
below or above a given range (using sr).
Methods
Channels that cannot be represented at the target EDF record duration (because their sample count per record would not be an integer) are dropped from the in-memory recording prior to any record-size restructuring. Optionally, channels outside a specified sample-rate range are also removed, preventing inadvertent inclusion of auxiliary low-rate channels in downstream analyses.
Parameters
| Parameter | Example | Description |
|---|---|---|
dur |
dur=1 |
Proposed EDF record size (seconds), which may differ from the current value |
sr |
sr=100,200 |
Optionally, drop signals with rates below 100 Hz or above 200 Hz |
Output
No output other than a message to the log (and altering the in-memory signal).
REFERENCE
Re-references signals with respect to one or more other signals
Methods
EEG signals are re-referenced by subtracting a reference signal (or the average of multiple reference signals) from each target channel. When a single reference channel is specified, this corresponds to a bipolar or linked-electrode reference; when multiple reference channels are provided, their arithmetic mean is subtracted, approximating a linked-mastoid or common-average-like reference restricted to the specified electrodes. A pairwise referencing mode applies distinct reference channels to corresponding target channels rather than averaging across all references. Optionally, all channels can be first resampled to a common rate before referencing to ensure sample-by-sample alignment.
Parameters
| Parameter | Example | Description |
|---|---|---|
sig |
sig=C3,C4 |
Signal(s) to re-reference |
ref |
ref=A1,A2 |
Signal(s) to provide the reference |
pairwise |
Perform pairwise re-referencing between sig and ref (see below) |
|
new |
new=C3_LM |
Generate a new channel instead of altering sig |
sr |
sr=128 |
Resample all channels first to this sample rate |
Both sig and ref are required parameters. If more than one
channel is given as the reference (in a comma-delimited list), the
average of those channels is used as the reference value. Alternatively, if the pairwise
option is also specified, then sig and ref should be of the same length. In this case,
rather than average multiple ref channels, the command would do a pairwise referencing:
Given the above sig and ref values in the table above: without the pairwise option, the
two channels would be calculated as:
--> C3 - (A1+A2)/2
C4 - (A1+A2)/2
pairwise:
---> C3 - A1
C4 - A2
If using new, only a single channel can be re-referenced.
Output
No output, other than a note to the log. In memory, the updated
sig channels will contain the re-referenced values.
DEREFERENCE
De-references signals with respect to one or more other signals
This command is a mirror of REFERENCE: instead of subtracting another reference signal, this simply adds it back in, i.e.
effectively removing an existing reference. The options (new, pairwise, sr, sig and ref) are otherwise similar,
see above for details.
Methods
A previously applied reference is reversed by adding back the reference signal to the target channel, recovering an approximation to the original unreferenced signal. This is the arithmetic inverse of the re-referencing operation.
MINMAX
Set digital and physical minimum and maximum values in the EDF header to be equal across multiple channels. This can be necessary to enable other software to be able to work with an EDF, by making it better conform to the EDF specification. Signals specified here must be comparable, e.g. all EEG and EOG with a common amplifier and ADC, and so are expected to have similar scaling and sensitivity (unit/bit) in the EDF.
Methods
The physical minimum and maximum values in the EDF header are harmonized across a set of comparable channels by setting all channels to the common extremes of the group. This operation adjusts the scaling factor (sensitivity, in units per bit) uniformly, ensuring consistent amplitude representation and EDF-specification compliance without altering the underlying signal values.
Parameters
| Parameter | Example | Description |
|---|---|---|
sig |
sig=C3,C4 |
Signals (two or more) to set group min/max values |
Output
No formal output is given. The channels are rescaled internally. Any
subsequent commands (i.e. including WRITE to write a new EDF) will
therefore be based on these new header values.
Example
Here we have an EDF with channels C3, C4, F3, F4, O1, O2, A1 and A2. The HEADERS
command shows that the channels have different physical min/max values, and therefore
different SENS values (scaling of micro-volts per bit in the EDF):
luna id01.edf -o out.db -s HEADERS
destrat out.db +HEADERS -r CH
ID CH DMAX DMIN PDIM PMAX PMIN SENS SR TYPE
id01.edf F3 32767 -32768 uV 1574.8 -3276.8 0.07403 500 EEG
id01.edf F4 32767 -32768 uV 1720.9 -3276.8 0.07626 500 EEG
id01.edf C3 32767 -32768 uV 3276.7 -3276.8 0.1 500 EEG
id01.edf C4 32767 -32768 uV 2034.5 -3276.8 0.08104 500 EEG
id01.edf O1 32767 -32768 uV 3276.7 -3276.8 0.1 500 EEG
id01.edf O2 32767 -32768 uV 3276.7 -3276.8 0.1 500 EEG
id01.edf A1 32767 -32768 uV 3238.6 -3276.8 0.09941 500 EEG
id01.edf A2 32767 -32768 uV 3255.3 -3276.8 0.09967 500 EEG
After running the MINMAX command, we see that the SENS values are now
set to be equal across all channels. This command will not fundamentally
change the underlying signal data, only the scaling in the EDF header.
luna id01.edf -o out.db -s 'MINMAX & HEADERS'
ID CH DMAX DMIN PDIM PMAX PMIN SENS SR TYPE
id01.edf F3 32767 -32768 uV 3276.7 -3276.8 0.1 500 EEG
id01.edf F4 32767 -32768 uV 3276.7 -3276.8 0.1 500 EEG
id01.edf C3 32767 -32768 uV 3276.7 -3276.8 0.1 500 EEG
id01.edf C4 32767 -32768 uV 3276.7 -3276.8 0.1 500 EEG
id01.edf O1 32767 -32768 uV 3276.7 -3276.8 0.1 500 EEG
id01.edf O2 32767 -32768 uV 3276.7 -3276.8 0.1 500 EEG
id01.edf A1 32767 -32768 uV 3276.7 -3276.8 0.1 500 EEG
id01.edf A2 32767 -32768 uV 3276.7 -3276.8 0.1 500 EEG
Note, if the EDF contained other signals that you did not want
included in the MINMAX procedure (e.g. respiratory channels, which
have different scaling from EEG channels), you would need to add sig
after MINMAX to specify, e.g. only the EEG channels. This command
will skip any EDF+ Annotation channels automatically.
uV
Converts a signal to uV units
Checks the unit (physical dimension) field of the EDF header for either V, mV or uV
and rescales the signal appropriately. If the header specifies some
other unit, or none, then no action is taken.
Methods
Signal amplitudes are converted to microvolts by reading the physical dimension field of the EDF header and applying the appropriate multiplicative scaling factor (×1,000 from mV, or ×1,000,000 from V). If the header dimension is already µV, or is unrecognized, no rescaling is applied.
Parameters
| Parameter | Example | Description |
|---|---|---|
sig |
sig=C3,C4 |
Signal(s) to convert |
If sig is not specified, this command is applied to all channels.
Output
No output, other than updating the in-memory signal.
mV
Converts a signal to mV units
Checks the unit (physical dimension) field of the EDF header for either V, mV or uV
and rescales the signal appropriately. If the header specifies some
other unit, or none, then no action is taken.
Methods
Signal amplitudes are converted to millivolts by reading the physical dimension field of the EDF header and applying the appropriate multiplicative scaling factor (÷1,000 from µV, or ×1,000 from V). If the header dimension is already mV, or is unrecognized, no rescaling is applied.
Parameters
| Parameter | Example | Description |
|---|---|---|
sig |
sig=C3,C4 |
Signal(s) to convert |
If sig is not specified, this command is applied to all channels.
Output
No output, other than updating the in-memory signal.
TIME-TRACK
Adds a time-track, which implicitly converts an EDF into an EDF+
This command is only used internally, currently.
Methods
The TIME-TRACK command converts a standard EDF to EDF+C format by appending an EDF Annotations channel that encodes the onset time of each record as a Time-stamped Annotation List (TAL). If the in-memory EDF is not already marked as EDF+, the header reserved field is updated accordingly and the file is declared continuous (EDF+C). A new annotation signal channel is then inserted at the end of each record, with a fixed width (controlled by globals::edf_timetrack_size), containing a TAL entry of the form +<onset>\x14\x14\x00 where <onset> is the elapsed time in seconds from the EDF start. For continuous recordings this offset is computed incrementally as a multiple of the record duration; for discontinuous (EDF+D) recordings produced by a merge operation, per-record timestamps are provided explicitly. The channel is labelled EDF Annotations (or EDF Annotations1, EDF Annotations2, etc. if annotation channels already exist), with nominal digital range −32768 to 32767 and scaling fields set to unity, as required by the EDF+ specification.
FLIP
Flips the polarity of a signal
Multiplies every sample value of a signal by -1.
Methods
Signal polarity is inverted by multiplying all sample values by −1, correcting for inadvertent polarity reversals introduced during recording or electrode placement.
Parameters
| Parameter | Example | Description |
|---|---|---|
sig |
sig=C3,C4 |
Signals to flip |
Output
No output, other than a message to the log and an updated in-memory signal.
Example
This next command takes the first 10 epochs of the C3 signal,
outputs the original signal to a file (f1), then flips the signal,
and re-outputs it (to f2):
luna me.lst sig=C3 -s 'EPOCH & MASK epoch=1-10 & \
RESTRUCTURE & MATRIX file=f1 & \
FLIP & MATRIX file=f2'
Note
In the above, we used the end-of-line \ character (with no trailing whitespace)
to continue the command on multiple lines, as many shells allow.
Comparing the original signals (looking at just the first 10 rows of output) ...
head f1
ID E S SP T C3
id001 1 0 0 0 3.43407
id001 1 0 1 0.00390625 2.06044
id001 1 0 2 0.0078125 -0.0763126
id001 1 0 3 0.0117188 -1.60256
id001 1 0 4 0.015625 -2.21306
id001 1 0 5 0.0195312 -2.21306
id001 1 0 6 0.0234375 -2.21306
id001 1 0 7 0.0273438 -2.06044
id001 1 0 8 0.03125 -2.06044
... to the new signals, we see the values have been flipped, albeit not as exactly as one might expect. (This is due to the encoding used by EDFs; see the note below for more details).
head f2
ID E S SP T C3
id001 1 0 0 0 -3.43865
id001 1 0 1 0.00390625 -2.06244
id001 1 0 2 0.0078125 0.0728122
id001 1 0 3 0.0117188 1.59799
id001 1 0 4 0.015625 2.20806
id001 1 0 5 0.0195312 2.20806
id001 1 0 6 0.0234375 2.20806
id001 1 0 7 0.0273438 2.05909
id001 1 0 8 0.03125 2.05909
Floating point accuracy
EDFs store data as 2-byte
integers: in contrast, floating point numbers as used
in Luna typically take up 4 or 8 bytes in memory. This relatively
low resolution of EDF introduces slight numerical differences so
that the values are clearly different from -1 times the original:
i.e. 3.43407 is not minus -3.43865. As noted in the EDF spec,
practically this limit on resolution is not a real issue for most
biosignals, if they are recorded with sensible physical and
digital min/max values to reflect the dynamic range of the signal.
ZC
Mean-center a signal
Subtracts the mean from a signal, either based on the entire duration or performed epoch-by-epoch. The latter may be more appropriate if there are large changes in the scale/mean of the signal across the recording.
Methods
Signals are mean-centered by subtracting either the whole-recording mean or, when epoch-wise centering is requested, the within-epoch mean from each sample. Epoch-wise centering removes slowly drifting DC offsets that vary across the night and may otherwise bias spectral or amplitude-based analyses.
Parameters
| Parameter | Example | Description |
|---|---|---|
sig |
C3,C4 |
Signals to flip |
epoch |
Perform mean-centering epoch-by-epoch |
Output
No output, other than a message to the log and an updated in-memory signal.
Example
Consider the SpO2 signal,
luna s.lst -s 'STATS sig=SpO2'
MEAN 94.1955
First adding the ZC command:
luna s.lst -s 'ZC sig=SpO2 & STATS sig=SpO2'
MEAN -0.000734802
Note that, due to EDF's 16-bit floating-point accuracy, the mean will
not be numerically exactly 0.00 (i.e. as internally, the signal is
written back to the internal, in-memory EDF at the end of the ZC
command, before being re-read by STATS).
ROBUST-NORM
Standardizes a signal using a robust approach
Normalizes a signal, using as measures of central tendency and spread the median and an estimate of the SD based on the inter-quartile range ( 0.7413 times IQR ). Additionally, this command can winsorize a signal (and optionally re-normalize after winsorization, to ensure (non-robust) mean/SD of 0/1). This can be performed either on the whole signal, or epoch-by-epoch.
Methods
Signals are standardized using a robust normalization procedure that subtracts the median and divides by a robust estimate of the standard deviation derived from the interquartile range (IQR × 0.7413, which equals the SD for a normal distribution). Because this approach uses rank-based statistics, it is resistant to the influence of extreme values and non-Gaussian tails that are common in physiological signals. Optionally, signal values beyond specified percentile thresholds can be winsorized (clipped to those quantile boundaries) prior to normalization, further limiting the influence of outliers. A second normalization pass may subsequently be applied to the winsorized signal to yield a distribution with mean 0 and SD 1. Normalization can be performed either across the whole signal or independently within each epoch, the latter being appropriate when the signal's scale changes systematically across the recording.
Parameters
| Parameter | Example | Description |
|---|---|---|
sig |
C3,C4 |
Signals to flip |
epoch |
Perform mean-centering epoch-by-epoch | |
center |
'F' | Perform median-centering (default: true ) |
scale |
'F' | Perform rescaling (default: true ) |
winsor |
0.05 | Winsorize the signal, e.g at 5th percentile (default: none) |
second-norm |
Perform a second round of normalization after winsorization (default: no) |
Output
No output, other than a message to the log and an updated in-memory signal.
Example
Consider the SpO2 signal,
luna s.lst -s 'STATS sig=SpO2'
MEAN 94.1955
SKEW -8.02593
If, for some reason, one wanted to normalize this measure, we can use ROBUST-NORM, also adding winsor=0.05 to
winsorize the signal at the 5th (and 95th) percentiles:
luna s.lst -s 'ROBUST-NORM sig=SpO2 winsor=0.05 & STATS sig=SpO2'
From the subsequent STATS output, we can see the mean and SD are closer to 0 and 1 respectively, and (due to the winsorization), the
signal is also much less skewed:
MEAN -0.80652
SKEW -0.4575
SD 2.05714
STATS command, we see the median (P50) is effectively 0 (save for numerical rounding):
P01 -5.39593
P02 -5.39593
P05 -5.39593
P10 -4.04687
P20 -2.69795
P30 -1.3489
P40 -1.3489
P50 -9.67455e-05
P60 -9.67455e-05
P70 -9.67455e-05
P80 1.48381
P90 1.48381
P95 2.83286
P98 2.83286
P99 2.83286
P01, P02 and P05 are all identical now.
If for some reason it is important to further rescale the signal to have mean and SD of 0 and 1 more precisely, then add the option second-norm to ROBUST-NORM. This results in the following mean and SD:
MEAN -5.44263e-06
SD 0.999983
SCALE
Rescale (and/or clip) a channel based on min-max ranges
If clip-min and/or clip-max is set, the signal will be clipped/winsorized at these thresholds (e.g. values
beyond those thresholds will be set to those values).
Additionally, if min-max is set to specify a lower/upper bound, the
scale will be rescaled to set the min/max at those values.
Methods
Signals are linearly rescaled to a specified output range by applying a min–max normalization. Optionally, values beyond specified lower and/or upper bounds are first clipped (winsorized) to those boundary values before rescaling, preventing extreme outliers from dominating the rescaled range.
Parameters
| Parameter | Example | Description |
|---|---|---|
min-max |
0,100 | Lower and upper bounds |
clip-min |
0 | Lower bound to clip (on original scale) |
clip-max |
1 | Upper bound to clip (on original scale) |
Output
No output other than changing signals in the internal EDF.
Example
The following assumes a signal (e.g. oxygen saturation) that is scored on a 0 - 1 scale, but contains outliers (below 0, or above 1, e.g. due to resampling or other artifact). The command below first clips at 0 and 1; it then rescales to a percent (0-100) rather than a proportion (0-1) scale:
luna s.lst -s ' SCALE sig=oxy clip-min=0 clip-max=1 min-max=0,100 '
Typically a SCALE command will be paired with subsequent analyses,
or a WRITE command to output a new EDF. (i.e. by itself the above command
would only change the in-memory EDF and then quit/move to the next EDF).
LOG
Apply a log transform to one or more signals
The LOG command applies a natural log transform to one or more signals:
log( x + offset )
This is useful when working with strongly right-skewed non-negative signals, for example activity counts, power-like measures, or other strictly positive derived channels.
Values below zero are clamped to zero before the transform, with a warning.
Methods
Signals are transformed in-place using the natural logarithm after the addition of a small constant offset. This offset can be fixed explicitly, which is often appropriate for count or activity data, or estimated from the observed signal distribution using a low percentile, which is often more appropriate for strictly positive continuous measures such as spectral power.
Parameters
| Parameter | Example | Description |
|---|---|---|
sig |
sig=activity |
One or more signals to transform |
offset |
offset=1 |
Fixed additive constant before taking the log |
eps |
eps=0.01 |
Data-driven offset: 0.1 times this percentile of the signal |
If both offset and eps are specified, offset takes priority.
For eps, Luna sets:
offset = 0.1 x percentile(sig, eps)
with a fallback to a very small constant if that percentile is zero or non-finite.
Output
No formal output other than changing the selected signals in the internal EDF and writing notes to the log.
Examples
Apply a standard log(x+1) transform to an actigraphy channel:
luna s.lst -s ' LOG sig=activity offset=1 '
Apply a percentile-based offset to a strictly positive derived signal:
luna s.lst -s ' LOG sig=DELTA eps=0.01 '
CLIP
Clip a signal using absolute or relative thresholds
The CLIP command truncates one or more signals in the in-memory EDF
so that values outside specified bounds are clipped to those bounds.
Unlike SCALE, which can optionally clip as part of a
rescaling step, CLIP is a standalone command intended specifically
for threshold-based clipping.
Thresholds can be specified either directly on the signal scale or via
quantiles of the observed signal distribution. This command modifies
the in-memory signal only; pair it with WRITE if
you want to save the clipped signal to a new EDF.
Methods
Signal amplitude is bounded (winsorized) by setting values below a specified lower threshold and/or above a specified upper threshold to those threshold values. Thresholds may be defined as absolute values on the native signal scale or as quantiles of the observed signal distribution, the latter providing a data-adaptive approach that automatically accommodates signals with different amplitude ranges.
Parameters
| Parameter | Example | Description |
|---|---|---|
sig |
sig=C3,C4 |
One or more signals to clip |
lwr |
lwr=-500 |
Absolute lower bound |
upr |
upr=500 |
Absolute upper bound |
lwr-pct |
lwr-pct=0.01 |
Lower clipping threshold set to this percentile |
upr-pct |
upr-pct=0.99 |
Upper clipping threshold set to this percentile |
Percentile thresholds must lie strictly between 0 and 1.
If multiple threshold options are given, Luna applies them in the following order for each selected signal:
- absolute lower bound (
lwr) - absolute upper bound (
upr) - percentile lower bound (
lwr-pct) - percentile upper bound (
upr-pct)
Output
No formal output other than changing the selected signals in the internal EDF and writing a note to the log.
Examples
Clip a signal to an absolute range:
luna s.lst -s ' CLIP sig=EMG lwr=-200 upr=200 '
Clip a signal based on distribution tails:
luna s.lst -s ' CLIP sig=C3 lwr-pct=0.01 upr-pct=0.99 '
As above, CLIP is distinct from the CLIP summary metric reported
by SIGSTATS, which quantifies the
proportion of clipped sample points rather than modifying the signal.
COMBINE
Combine two or more channels to create a new channel
This command allows new channels to be created based on simple sum/mean/median of one or more channels.
By default, this command requires that all channels requested (by sig) are present; an
error will be given otherwise. This behavior can be changed by adding allow-missing,
which means that only the available channels are used (e.g. if only 2 of 3 found, the
sum would be based just on those two). If no channels are available, no new
channel is created. If only two channels are available and a median is requested, this is
automatically changed to a mean instead.
Methods
A new channel is constructed as the sample-wise sum, arithmetic mean, or median of two or more existing channels. This operation is used, for example, to create a composite respiratory effort channel, a bilateral EEG average, or to combine reference electrodes prior to referencing.
Parameters
| Parameter | Example | Description |
|---|---|---|
sig |
C3,C4 |
Signal list |
sum |
S1 |
Make a new sum channel called S1 |
mean |
M1 |
Make a new mean channel called M1 |
median |
M1 |
Make a new median channel called M1 (for 3 or more channels) |
allow-missing |
If present, allow for channels in sig to be not present |
Names for new channels (e.g. S1 or M1 above) are not allowed to exist in the EDF prior to calling COMBINE.
Output
No output other than adding new signals to the internal EDF.
Example
To create a new channel S that is the sum of two channels ABDO and
THOR, for example, and then output epoch-level statistics for this
new channel:
luna s.lst -s ' COMBINE sum=S sig=THOR,ABDO
STATS sig=S epoch '
RECORD-SIZE
Alters the record size of an EDF
This command changes the low-level encoding of data in an EDF, which is something that you should not normally need to change. Often, EDFs have a record size (i.e. the size of the blocks in which the data are stored) of 1 second or so. Why might you want to change this?
-
as the smallest
EPOCHsize is limited by the EDF record size, if the EDF record size is relatively large (e.g. 10 seconds), it will not be possible to specify smaller epochs (e.g. 5 seconds). -
if the EDF record size is very small (e.g. 100 milliseconds), this can reduce performance when reading the EDF from disk
There are a number of points that should be borne in mind:
-
no subsequent commands can be issued after a
RECORD-SIZEcommand; rather, a new EDF will be written to disk -
you should ensure that the new record size contains an integer number of samples for all signals
-
currently, you can only change the record size of EDF, not EDF+ files
-
as only whole records are written to disk, the final part of an EDF (that is shorter than the new record size) may be truncated
Methods
The low-level record (block) size of the EDF is changed by reorganizing samples into records of the specified duration. This operation is required when the original record size is incompatible with a desired epoch length or imposes an unnecessary read-performance penalty. The modified EDF is immediately written to disk; any trailing samples that do not fill a complete record at the new block size are discarded.
Parameters
| Parameter | Example | Description |
|---|---|---|
dur |
dur=1 |
New EDF record/block size |
edf-dir |
edf-dir=edfs/ |
Folder for writing new EDFs |
edf-tag |
edf-tag=rec1 |
Tag added to new EDFs |
sample-list |
sample-list=s2.lst |
Generate a sample-list pointing to the new EDFs |
That is, while RECORD-SIZE itself only takes dur as the single
option, one must also specify all options for
WRITE, as RECORD-SIZE automatically triggers
WRITE after changing the record size of the in-memory
representation. (That is, as always, the original EDF file is left
untouched.)
Output
No output, other than message to the log and an updated in-memory signal.
Example
Focusing only on the signals PR and EEG in the first
tutorial EDF, we see that this EDF has a record size
of 1 second:
luna s.lst 1 sig=PR,EEG -s 'SUMMARY'
# signals : 2
# records : 40920
Duration : 1
That is, the EDF has 40,920 records, each of duration 1 second.
Looking at the two signals, because the record duration is 1 second,
this implies a sample rate of 1 Hz and 125 Hz respectively for PR
and EEG.
Signal 1 : [PR]
# samples per record : 1
...
Signal 2 : [EEG]
# samples per record : 125
...
To generate a new EDF (which contains only these two signals) with an altered record size (in this example, 50 seconds):
luna s.lst 1 sig=PR,EEG -s 'RECORD-SIZE dur=50 edf-tag=r50'
(Note that setting a 50-second record size would be unusual, this is done here purely for illustrative purposes.) After running this command, you'll see the following messages in the log:
saved new EDF, edfs/learn-nsrr01-r50.edf
**warning: the PROBLEM flag was set, skipping to next EDF...
The warning message is expected, this is just Luna's way of ensuring
that no further commands can be run after RECORD-SIZE command.
Running SUMMARY on the new EDF, we see that the record size has been
changed:
luna edfs/learn-nsrr01-r50.edf -s SUMMARY
# records : 818
Duration : 50
...
Signal 1 : [PR]
# samples per record : 50
...
Signal 2 : [EEG]
# samples per record : 6250
...
That is, instead of 40,920 records of 1 second we have 818 records of 50 seconds. Correspondingly, there are now 50 times the number of samples per record compared to the original EDF (the sample rate in Hz is obviously the same as before).
Note
Because 40,920 is not evenly divisible by 50, the last 20
seconds has been truncated (i.e. the log will indicate a total
duration of 11:21:40 instead of the original 11:22:00).
EDF-MINUS
Collapsed gapped EDF+D records to standard, aligned EDFs
EDF+D files are convenient for recording data that have gaps. However, it can sometimes be inconvenient to analyse recordings with gaps, and some other tools do not accept EDF+D files. When the goal is to convert a gapped recording and output a new EDF, the record-based structure of EDF can be a problem: whereas gaps can be fractions of an EDF record (e.g. often but not always 1 second), an EDF file must contain only whole records.
More generally, a standard EDF with an "ideal" structure might be as follows:
-
data in a single segment
-
one-to-one correspondence between stage annotations and epochs (units of analysis)
-
a clean mapping of EDF records to annotations: stages are a fixed integer number of seconds and annotations and epochs align to EDF record boundaries
In practice, many records (especially from clinical studies) are EDF+D with less-than-ideal properties from the perspective of clean and convenient analysis:
-
discontinuous segments (gaps); often only a single (largest) segment of interest, but not always
-
stage annotations not regularly aligned across the night and/or with standard epochs
-
stage annotations not aligned with EDF record boundaries
-
potentially, EDF records are of nonstandard duration (e.g. 3.92 seconds) or very long (e.g. 30 seconds)

In terms of analysis:
-
Luna naturally represents gapped EDF+ -- in fact, internally, after any type of markings (e.g. restriction to NREM) the data structure is implicitly identical to an EDF+D file
-
epochs (unit of analysis) can be defined to align with stage annotations within each segment (
EPOCH align), even ones of variable duration via generic epochs (EPOCH annot) -
however, some analyses (e.g. summarizing hypnogram structures/timings) can be challenging with gapped recordings & incomplete annotations, or presence of variably-sized or incomplete epochs, however
In terms of file (EDF) generation:
-
when writing a new EDF, the atomic EDF record structure comes into play: we can only read or write whole records
-
also, some EDFs have awkward record sizes (e.g. 3.92 seconds, and in some cases may not even allow an integer number of evenly-spaced samples)
-
in PSGs containing signals of variable sampling rate, we cannot arbitrarily change record size (i.e. each record must contain a fixed, positive integer number of samples for every signal)
-
if stage annotations & record boundaries do not align, we cannot easily output a new EDF that excludes/includes only a given stage
-
when trying to align signals & annotations, timing difficulties arise due to EDF+ varying its specification of time:
1) EDF start time constrained to whole seconds (hh:mm:ss),
2) signal segments constrained to whole record units,
3) no constraints on annotation start/stops,
4) no constraints on gap durations
Further, when Luna masks a recording (by flagging epochs as masked or not), the actual transformation is at the level of EDF records. Here, records and epochs align, such that if we mask epoch 2, this cleanly splices out records 3 and 4:
Record 1--|2--|3--|4--|5--|6--|7--|8--|
Epoch 1------|2------|3------|4-------
Masked epochs XXXXXXXX
Retained records 1--|2--| 5--|6--|7--|8--|
In contrast, here records and epochs do not align: Luna will retain (whole) records that have at least some unmasked signal, so here, only record 4 would be dropped:
Record 1--|2--|3--|4--|5--|6--|7--|8--|
Epoch 1------|2------|3------|4------|
Masked epochs XXXXXXXX
Retained records 1--|2--|3--| 5--|6--|7--|8--|
Obviously, in the context of analysis (or exporting in formats other
than EDF), it is possible to slice data at a sample-point resolution
(and, if necessary, to resample different signals to have similar
sample rates, further simplifying things). However, if we essentially want to
retain the native EDF (record-based) properties, then it can be useful to manipulate
the EDF structure to streamline downstream analysis, which is where EDF-MINUS comes in.
Specifically, EDF-MINUS is a command to make standard EDFs with more convenient record/annotation/epoch structure:
-
sets EDF record size to a fixed 1 second
-
selects and/or combines gapped segments (e.g. all, largest, only those w/ staging information, etc)
-
either splices out gaps or zero-pads to fill gaps, but also ensuring all points align with EDF record boundaries
-
optionally aligns segments to stage annotation boundaries
-
shifts all annotation start/stops as needed (i.e. if splicing out gaps) to keep annot/signal alignment
-
adds annotations to show original (gapped) structure
-
saves new EDF and annotation files
When handling gaps, there are two choices:
-
keep timeline as is: fill the gaps in the signal (zero-padding) and leave annotations as they are
-
splice out gaps: this "ignores" gaps, similar to simply reading an EDF+D as a standard EDF, except here
EDF-MINUSa) still aligns retained segments to record and/or epoch/staging boundaries, and b) alter any annotation timelines to be locally-consistent with the new spliced set of signals
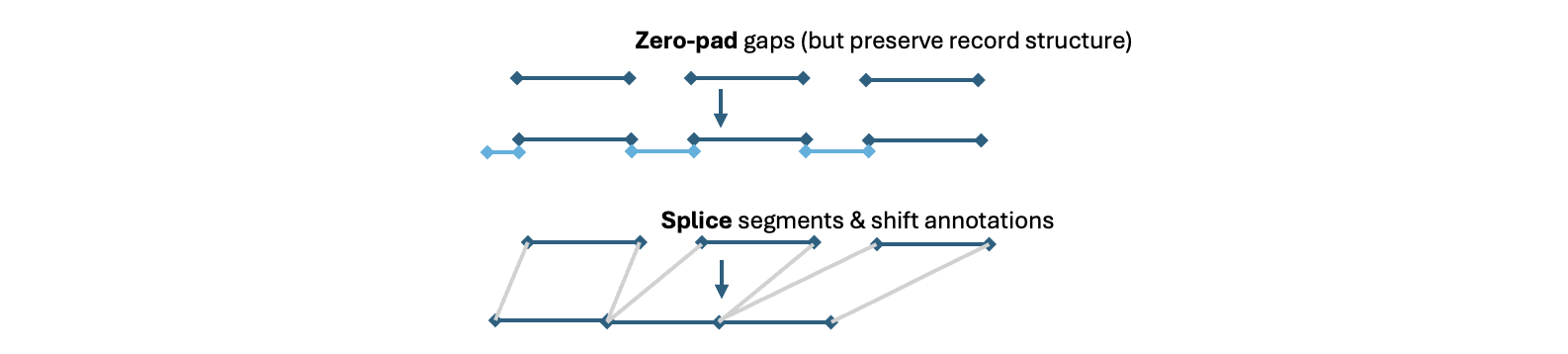
Which strategy should you choose? Practically, in the context of PSG/sleep data, it is usually sensible to pick what best aligns to the staging information. If staging is "constant" (with respect to clock-time, e.g. truly every 30 seconds), then it makes sense to keep that as is and zero-pad. Staging is not constant across different segments (i.e. as if each segment was scored separately), then it is better to splice out the gaps.
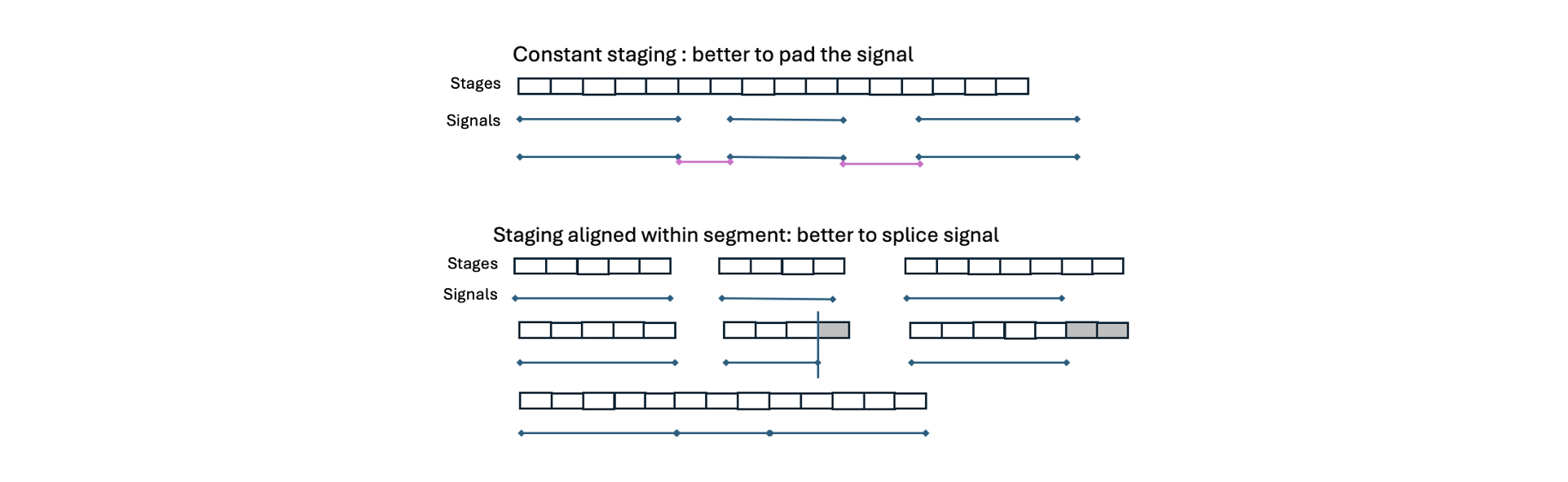
Obviously both these operations will slightly change the signal
data. If zero-padding, one would need to handle those regions in
downstream analysis, but EDF-MINUS provides an annotation to
indicate which regions are zero-padded (e.g. and can be more cleanly
extracted from analysis, as those regions will now also align with EDF
record/epoch boundaries). In most cases, one may drop a few seconds
of signal, etc, but given that signal is (by definition) dropped in
the context of a gapped recording, it is often a reasonable trade-off
to drop a little more data to make the resulting records easier to
work with.
Methods
Discontinuous EDF+D recordings are converted to standard, contiguous EDF files using a two-stage alignment procedure. First, the desired segments are selected (e.g., only the largest segment or those containing scoring annotations), and records are realigned to standard epoch and record boundaries using a sub-sample interpolation step. Gaps between segments are handled by one of two strategies: zero-padding, which preserves the original clock-time timeline by inserting silent intervals at gap locations and annotating them accordingly; or splicing, which concatenates retained segments directly, with annotation timestamps adjusted to reflect the new local timeline. In both cases, the output is a standard EDF (without discontinuities) in which record boundaries, epoch boundaries, and staging annotations are mutually aligned, facilitating downstream analysis and compatibility with EDF-only software.
Parameters
Main arguments
| Parameter | Example | Description |
|---|---|---|
policy |
splice |
Treatment of segments (splice, zero-pad ) |
segments |
2,3,4 | Numbers of segments to keep, or largest (default: keep all) |
out |
new1 |
Required name for new EDF/annotations |
require |
N1,N2,N3,R,W |
Only select segments with at least one of these annotations having any extent of overlap |
require-dur |
30 | Require at least N seconds to be spanned by required annotation(s) |
require-whole |
N1,N2,N3,R,W |
Alternative to require but insists that segments must span whole annotations |
align |
N1,N2,N3,R,W |
Align & truncate segments to these annotations (i.e. stages) |
dur |
30 | Annotation/epoch duration for alignment (e.g. assuming (multiples of) fixed 30s epochs by default) |
hms |
Output annotation files in h:m:s clock-time format (not elapsed seconds) | |
dhms |
Output annotation files in date+clock-time format |
The require options can be used to select only segments that have some extent of staging information.
Secondary arguments
| Parameter | Example | Description |
|---|---|---|
max-sr |
1024 | Optionally, set a warning if sample rate is above this value |
prefix |
edit1 |
Optional prefix for new book-keeping annotations added (e.g. tracking splice-points) (default: none) |
Note: EDF-MINUS applies a zero-hold filter on signals with <1 Hz
sample rate (must be 1/N Hz where N is an integer) to fit in
1-second EDF record
Output
The primary output is a new (standard) EDF file and optionally an annotation (.annot) file.
Example
Consider this example gapped EDF+D:
luna m.lst -s SEGMENTS
Processing: id01 [ #1 ]
duration 06.30.59, 23459s | time 23.19.59 - 05.51.25 | date 07.04.16
signals: 23 (of 23) selected in an EDF+D file
F3_M2 | F4_M1 | C3_M2 | C4_M1 | O1_M2 | O2_M1 | EOG_L | EOG_R
CHIN_1 | EKG | R_R | LEG_L | LEG_R | Snore | Airflow | PTAF
Chest | Abdomen | SUM | SAO2 | PLETH | ETCO2 | EDF Annotations
extracting 'EDF Annotations' track from EDF+
annotations:
? (x41) | N1 (x72) | N2 (x74) | W (x593)
edf_annot (x0) | note (x138)
Reviewing the location of segments:
destrat out.db +SEGMENTS -r SEG -v START_HMS STOP_HMS DUR_MIN
ID SEG DUR_MIN START_HMS STOP_HMS
id01 1 96.7166 23:19:59.000 00:56:42.000
id01 2 71.7666 00:56:49.000 02:08:35.000
id01 3 82.65 02:08:42.000 03:31:21.000
id01 4 103.766 03:31:27.000 05:15:13.000
id01 5 36.0833 05:15:20.000 05:51:25.000
As it contains staging information, we can extract that also epoch-by-epoch:
luna m.lst -o out.db -s STAGE
destrat out.db +STAGE -r E
ID E CLOCK_TIME START_SEC STAGE
id01 1 23:19:59 0 L
id01 2 23:20:29 30 L
id01 3 23:20:59 60 L
id01 4 23:21:29 90 L
id01 5 23:21:59 120 L
...
id01 194 00:56:49 5810 W
id01 195 00:57:19 5840 W
id01 196 00:57:49 5870 W
id01 197 00:58:19 5900 W
id01 198 00:58:49 5930 W
id01 199 00:59:19 5960 W
id01 200 00:59:49 5990 W
Note that the timing of the available staging changes across the night - e.g. every 59-th and 29-th second past the minute initially, but then at the 19/49-second marks later on. We can assume that within-segment staging is well aligned - i.e. we do not see any "conflict" messages in the output above.
To go from EDF+D to a standard EDF, there are two routes/policies:
- assume staging is fixed and correct (w.r.t. clock time) ->
zero-pad signals; this changes signals (fills gaps, to nearest
EDF record unit), keeps annotations (largely) 'as is'
- or, assume staging is not aligned uniformly w.r.t. clock time ->
splice signal; keeps signals largely 'as is' (i.e. ignores gaps,
implicitly sets to 0s dur) but rather change annotations to
align
In this example, the annotations are not cleanly and consistently aligned w.r.t. clock-time (e.g. always 0 and 30-secs past the minute); thus, we do not want to zero-pad (which keeps annotations largely as is).
So, we'd select 'splice' mode: by default, this uses the staging
annotations to align segments (N1, N2, N3, R, W and ?),
editing segments such that the resulting output is cleanly/consistently
aligned. We'll put the new EDF in the folder fin/:
mkdir fin
EDF-MINUS:
luna m.lst -o out.db -s EDF-MINUS out=fin/splice policy=splice
The console gives information on the option settings:
CMD #1: EDF-MINUS
options: out=fin/splice policy=splice sig=*
settings:
join-policy (policy) = splice
retained segments (segments) = all
maximum sample rate allowed (max-sr) = 1024 Hz
segment alignment annotations (align) = ?,N1,N2,N3,R,W
alignment duration unit (dur) = 30s
required annotations (require) =
require whole annots (require-whole) = F
require at least (require-dur) = 0s
annotation prefix (prefix) =
output file-root (out) = fin/splice
It then further lists the structure of the EDF+D
dataset contains 22 signals and 6 annotation classes (918 instances)
specified 6 annotation classes (?,N1,N2,N3,R,W) for alignment (780 instances found)
aligning segment 0.00->5803.00 start to 0 secs based on annotation ? = 0.00->30.00
& aligning segment end to 5790 based 193 whole intervals of 30s from aligned start at 0s
aligned segment 1 : 0.00-5803.00 --> 0.00-5790.00
aligning segment 5810.00->10116.00 start to 5810 secs based on annotation W = 5810.00->5840.00
& aligning segment end to 10100 based 143 whole intervals of 30s from aligned start at 5810s
aligned segment 2 : 5810.00-10116.00 --> 5810.00-10100.00
aligning segment 10123.00->15082.00 start to 10123 secs based on annotation W = 10123.00->10153.00
& aligning segment end to 15073 based 165 whole intervals of 30s from aligned start at 10123s
aligned segment 3 : 10123.00-15082.00 --> 10123.00-15073.00
aligning segment 15088.00->21314.00 start to 15088 secs based on annotation W = 15088.00->15118.00
& aligning segment end to 21298 based 207 whole intervals of 30s from aligned start at 15088s
aligned segment 4 : 15088.00-21314.00 --> 15088.00-21298.00
aligning segment 21321.00->23486.00 start to 21321 secs based on annotation W = 21321.00->21351.00
& aligning segment end to 23481 based 72 whole intervals of 30s from aligned start at 21321s
aligned segment 5 : 21321.00-23486.00 --> 21321.00-23481.00
The five segments are spliced and realigned:
found 5 segment(s)
[ original segments ] -> [ aligned, edited ] --> [ final segments ]
++ seg #1 : 0.00-5803.00 (5803s) [included] --> 0.00-5790.00 --> 0.00-5790.00 (13s shorter)
- gap #2 : 5803.00-5810.00 (7s) [spliced]
++ seg #2 : 5810.00-10116.00 (4306s) [included] --> 5810.00-10100.00 --> 5790.00-10080.00 (16s shorter)
- gap #3 : 10116.00-10123.00 (7s) [spliced]
++ seg #3 : 10123.00-15082.00 (4959s) [included] --> 10123.00-15073.00 --> 10080.00-15030.00 (9s shorter)
- gap #4 : 15082.00-15088.00 (6s) [spliced]
++ seg #4 : 15088.00-21314.00 (6226s) [included] --> 15088.00-21298.00 --> 15030.00-21240.00 (16s shorter)
- gap #5 : 21314.00-21321.00 (7s) [spliced]
++ seg #5 : 21321.00-23486.00 (2165s) [included] --> 21321.00-23481.00 --> 21240.00-23400.00 (5s shorter)
original total duration = 23459s
retained total duration = 23400s (59s shorter)
Finally, it writes the new EDF:
creating a new EDF fin/splice.edf with 22 channels
retaining original EDF start-time of 23.19.59
retaining original EDF start-date of 7.4.2016
created an empty EDF of duration 23400 seconds
creating annotation file fin/splice.annot with 914 annotations from 5 classes
data are not truly discontinuous
writing as a standard EDF
writing 22 channels
saved new EDF, fin/splice.edf
writing annotations (.annot format) to fin/splice.annot
We can check this new file:
luna --build fin > f.lst
splice fin/splice.edf fin/splice.annot
We'll use the SPANNING command to give some information on the new EDF, which also confirms this is a standard EDF:
luna f.lst -o out.db -s SPANNING annot=N1,N2,N3,R,W,?
duration 06.30.00, 23400s | time 23.19.59 - 05.49.59 | date 07.04.16
signals: 22 (of 22) selected in a standard EDF file
We see that there is a 1-to-1 matching between annotations (staging) and EDF epochs now, and alignment between epochs, stage annotations (and EDF records):
destrat out.db +SPANNING | behead
ID splice
ANNOT_HMS 06:30:00.000
ANNOT_N 780
ANNOT_OVERLAP NO
ANNOT_SEC 23400
INVALID_N 0
INVALID_SEC 0
NSEGS 1
REC_HMS 06:30:00.000
REC_SEC 23400
SPANNED_HMS 06:30:00.000
SPANNED_PCT 100
SPANNED_SEC 23400
UNSPANNED_HMS 00:00:00.000
UNSPANNED_PCT 0
UNSPANNED_SEC 0
VALID_N 780
If one wants clock times in the new .annot, add hms or dhms to
make the generated .annot file have those versus elapsed seconds.
If we instead were to zero pad the above EDF+D:
luna m.lst -o out.db -s EDF-MINUS out=fin/zero policy=zero-pad
Note - in this instance, we will still trim segments to align w/ starts & duration of staging; in this instance, zero-padding doesn't really make sense as annotations are not cleanly consistent anyway.
These again are the detected and aligned segments:
dataset contains 22 signals and 6 annotation classes (918 instances)
specified 6 annotation classes (?,N1,N2,N3,R,W) for alignment (780 instances found)
aligning segment 0.00->5803.00 start to 0 secs based on annotation ? = 0.00->30.00
& aligning segment end to 5790 based 193 whole intervals of 30s from aligned start at 0s
aligned segment 1 : 0.00-5803.00 --> 0.00-5790.00
aligning segment 5810.00->10116.00 start to 5810 secs based on annotation W = 5810.00->5840.00
& aligning segment end to 10100 based 143 whole intervals of 30s from aligned start at 5810s
aligned segment 2 : 5810.00-10116.00 --> 5810.00-10100.00
aligning segment 10123.00->15082.00 start to 10123 secs based on annotation W = 10123.00->10153.00
& aligning segment end to 15073 based 165 whole intervals of 30s from aligned start at 10123s
aligned segment 3 : 10123.00-15082.00 --> 10123.00-15073.00
aligning segment 15088.00->21314.00 start to 15088 secs based on annotation W = 15088.00->15118.00
& aligning segment end to 21298 based 207 whole intervals of 30s from aligned start at 15088s
aligned segment 4 : 15088.00-21314.00 --> 15088.00-21298.00
aligning segment 21321.00->23486.00 start to 21321 secs based on annotation W = 21321.00->21351.00
& aligning segment end to 23481 based 72 whole intervals of 30s from aligned start at 21321s
aligned segment 5 : 21321.00-23486.00 --> 21321.00-23481.00
This is the impact of zero-padding:
found 5 segment(s)
[ original segments ] --> [ aligned, edited final segments ]
++ seg #1 : 0.00-5803.00 (5803s) [included] --> 0.00-5790.00 (13s shorter)
- gap #2 : 5803.00-5810.00 (7s) [zero-padded] --> 5790.00-5810.00 (13s longer)
++ seg #2 : 5810.00-10116.00 (4306s) [included] --> 5810.00-10100.00 (16s shorter)
- gap #3 : 10116.00-10123.00 (7s) [zero-padded] --> 10100.00-10123.00 (16s longer)
++ seg #3 : 10123.00-15082.00 (4959s) [included] --> 10123.00-15073.00 (9s shorter)
- gap #4 : 15082.00-15088.00 (6s) [zero-padded] --> 15073.00-15088.00 (9s longer)
++ seg #4 : 15088.00-21314.00 (6226s) [included] --> 15088.00-21298.00 (16s shorter)
- gap #5 : 21314.00-21321.00 (7s) [zero-padded] --> 21298.00-21321.00 (16s longer)
++ seg #5 : 21321.00-23486.00 (2165s) [included] --> 21321.00-23481.00 (5s shorter)
original total duration = 23459s
retained total duration = 23427s (32s shorter)
However, because the annotations have not been changed, you'd get
conflict messages if attempting commands such as HYPNO, for example
(which could not be solved by EPOCH align, as the staging
annotations themselves are not uniformly distributed.
ANON
Sets the in memory EDF header fields Patient ID and Start Date
fields to missing values as per the EDF spec (e.g. X X X X for EDF+ files, . for EDF). Any output
of EDFs subsequently generated with the WRITE command will have those fields blanked.
As with all Luna commands, this does not alter the original EDF.
Also see the anon=T special variable,
which wipes EDF headers before attaching any annotation files.
Methods
Protected health information is removed from the in-memory EDF header by replacing the patient identifier and recording start-date fields with the EDF-specification placeholder values, producing de-identified output EDFs when subsequently written to disk.
Note
This command does not alter the ID specified in the
sample-list (i.e. the first column). That ID,
which is used to track all output, etc, is distinct from the EDF
header Patient ID field, and may or may not be similar.
Parameters
| Option | Example value | Description |
|---|---|---|
insert-id |
If specified, will set the Patient ID to the sample-list ID instead of null | |
root |
cohort |
Will sequentially set IDs to cohort_1, cohort_2, etc |
Output
No output other than message to the log, and altering the in-memory representation of the EDF header.
Example
A typical EDF with identifying information in the header (showing only
relevant rows from the SUMMARY output):
luna my.edf -s 'SUMMARY' | head
EDF filename : my.edf
Patient ID : id00001
Recording info :
Start date : 07.06.16
Start time : 23:07:56
... (cont'd) ...
Here we see how the ANON command effectively wipes this information:
luna my.edf -s 'ANON & SUMMARY' | head
EDF filename : my.edf
Patient ID : .
Recording info :
Start date : .
Start time : 23:07:56
This next command takes all EDFs in a project (defined by s.lst) and
creates a set of new EDFs with the WRITE command
(in the folder edfs/, and with the new sample list s2.lst) that
are identical except they have the Patient ID and Start Date fields
set to missing:
luna s.lst -s 'ANON & WRITE edf-dir=edfs/ edf-tag=anon sample-list=s2.lst'
SET-HEADERS
Directly specify certain EDF header values
Note that EDF header fields which relate to the size/structure of the
actual data (i.e. sample rate, EDF header size, physical min/max, etc)
cannot be changed in this way - i.e. for those changes, use the relevant
data-modifying command, e.g. RESAMPLE, RECORD-SIZE, MINMAX,
etc). That is, these options only modify the header and nothing else.
Note that any start time/date changes and made after any annotations
are attached. Internally, annotations are represented as elapsed time
from the current EDF start - i.e. their alignment with the signal data
will not change, but the output of WRITE-ANNOTS will differ if
setting hms or dhms flags. See also the
starttime and
startdate special variables
(which make the changes before attaching annotations).
Methods
Descriptive EDF header fields (patient identifier, recording information, start date and time, channel transducer type, physical dimension, and prefiltering description) are updated in-memory without altering the underlying signal data or its scaling. This is used to correct erroneous metadata, apply standardized labels, or prepare files for redistribution.
Parameters
Primary headers
| Parameter | Example | Description |
|---|---|---|
id |
id001 |
EDF patient ID (max 80 chars) |
recording-info |
EDF recording information field (max 80 chars) | |
start-date |
EDF start date (max 8 characters; no format imposed) | |
start-time |
EDF start time (max 8 characters; no format imposed) |
Channel-specific headers
| Parameter | Example | Description |
|---|---|---|
sig |
${eeg} |
Specify the channel(s) to modify |
transducer |
Set the transducer field for specified channels (max 80 chars) | |
physical-dimension |
Set the physical dimension (units) for specified channels (max 8 chars) | |
unit |
Same as physical-dimension |
|
prefiltering |
Set the prefiltering field for specified channels (max 80 chars) |
Output
This command only modifies the in-memory representation of the EDF. Use WRITE to save any changes.
e.g. to fix the start date, and add transducer types for the EEGs (here using Luna's automatic specification of common EEG channel names):
luna file1.edf -s 'SET-HEADERS start-date=08.08.21 sig=${eeg} transducer=Ag/AgCl & WRITE edf-tag=edit '
file1-edit.edf with the above fields fixed.
SET-VAR
Directly specify Luna variables
This can be useful if one wants to include individual-level variables when defining a script variable. If it isn't clear why you'd want to do this, then there is no need to worry about using this command.
Methods
Named variables are set or overridden within the active Luna session, allowing script-level parameters to be dynamically assigned during processing rather than requiring all values to be specified at launch.
Parameters
This command takes only a single option, which will be given as the variable name. The argument is the variable value.
| Parameter | Example | Description |
|---|---|---|
| any valid variable name | v=100 |
Sets this variable (e.g. v) to the value 100 |
Output
None, other than a message to the console log.
Example
Sets an individual-level variable var to the text string val:
luna s.lst -s 'SET-VAR var=val'
SET-TIMESTAMPS
Directly specify EDF record timestamps
This is an advanced function for directly manipulating EDF record timing. Given a file with as many rows/values as there are EDF records in the current in-memory EDF, set each record to start at that time. This can be used to generate toy datasets, e.g. with gaps. Note that this command does not adjust annotations in any way.
All times are expected in seconds, one value per line, and all values must be increasing.
Methods
EDF record timestamps are directly overwritten using values supplied from an external file, enabling construction of recordings with arbitrary temporal structures (e.g., discontinuous segments) for testing or simulation purposes. Annotation timings are not adjusted by this operation.
Parameters
| Parameter | Example | Description |
|---|---|---|
file |
timestxt |
Required text file of new timestamps |
Output
None, except modifying the in-memory EDF and some information to the console.
Example
See this vignette for an example of using SET-TIMESTAMPS to generate an example EDF+D file.
RECTIFY
Rectifies a signal
This command sets all values of an EDF signal to their absolute values. It is primarily designed for use
working with other functions such as HILBERT and PEAKS, to build up larger processing procedures.
Methods
Full-wave rectification replaces each sample with its absolute value, collapsing the signal onto the non-negative real axis. This operation is commonly applied as a preprocessing step prior to envelope extraction or peak detection, where the sign of the signal is uninformative.
Parameters
| Parameter | Example | Description |
|---|---|---|
sig |
C3,C4 |
Signals to be rectified (or all, if this is absent) |
Output
None.
Example
As a toy example, here we rectify a signal with positive and negative values (an EEG):
luna s.lst 1 -o out.db \
-s ' TAG run/1 & STATS sig=EEG
& RECTIFY sig=EEG
& TAG run/2 & STATS sig=EEG '
destrat out.db +STATS -r CH run/1 -v MIN MAX | behead
ID nsrr01
CH EEG
run 1
MAX 125
MIN -124.019607843137
destrat out.db +STATS -r CH run/2 -v MIN MAX | behead
ID nsrr01
CH EEG
run 2
MAX 125
MIN 0.490196078431372
REVERSE
Reverse a signal
This command is primarily designed for evaluating other time-domain/phase-based methods, e.g. to provide a sanity-check by completely reversing a signal in the time-domain.
Methods
Time-domain reversal reflects the signal about its midpoint, so that the sample originally at time t is placed at time T − t, where T is the total recording duration. Because this transformation preserves the amplitude spectrum while inverting the phase spectrum, it serves as a useful negative control for assessing directional or causal properties of time-domain analyses.
Parameters
| Parameter | Example | Description |
|---|---|---|
sig |
C3,C4 |
Signals to be reversed (or all, if this is absent) |
Output
None (other than reversing the in-memory signal).
MOVING-AVERAGE
Applies a moving-average (or median) filter to a signal
Applies a moving average window to filter a signal based on either a) the mean, b) median, or c) ...
Methods
A sliding window of user-specified duration is applied to each signal to produce a smoothed output. Within each window, the central sample is replaced by the arithmetic mean, the median, or — for a triangular filter — a linearly weighted mean that tapers toward the window edges. Mean filtering attenuates high-frequency noise while preserving low-frequency trends; median filtering is more robust to transient outliers; and the triangular (Bartlett) weighting provides a smoother spectral roll-off than a rectangular window while remaining computationally efficient.
Parameters
Note that median and tri cannot be specified together.
| Parameter | Example | Description |
|---|---|---|
sig |
C3,C4 |
Signals to be reversed (or all, if this is absent) |
median |
Signals to be reversed (or all, if this is absent) | |
tri |
Signals to be reversed (or all, if this is absent) | |
hw |
Half-width (in seconds) of the triangular window | |
lwr |
Weight at triangular window edge (between 0 and 1) | |
epoch |
Signals to be reversed (or all, if this is absent) |
Output
None.
INSERT
Estimate lag between EDFs or insert channels from one EDF into another
INSERT supports two related workflows and is designed for
aligning and splicing channels across EDFs:
- Estimate the temporal offset between a primary and secondary EDF.
- Insert one or more channels from the secondary EDF into the current EDF using a known offset, optionally with a simple linear time-stretch correction.
Unlike most EDF-manipulation commands, insert mode can add channels at the secondary signal's own sample rate, so it is intended for merging asynchronous or differently sampled recordings into one in-memory EDF.
Current implementation notes:
- Both EDFs must be continuous, or at least not actually discontinuous with gaps.
- Lag-estimation mode requires matched sample rates within each comparison pair, and all comparison pairs must share the same sample rate.
- Insert mode zero-pads uncovered regions and makes channel names unique by
appending
.1,.2, etc. if needed. - The
annot=argument is parsed in insert mode, but annotation creation is not currently implemented.
Methods
INSERT now uses cross-correlation as the standard
alignment method. In estimation mode, matched windows are taken from the secondary
EDF, optionally band-pass filtered, z-scored, and aligned to the primary EDF by
cross-correlation. The per-pair lag estimates are combined by taking the median
lag across signal pairs for each window. Each window is then quality-gated using
the median matched-window correlation peak (min-peak, default 0.3), and only
accepted windows enter the drift model.
Across accepted windows, ordinary least-squares regression of estimated waveform
shift (seconds) on window time (seconds) is performed. Windows with residuals
greater than 3 SD are removed and the regression is refit. The fitted intercept
is the start-of-record waveform shift, and the fitted slope quantifies linear
clock drift between devices in seconds per second. When valid EDF start times are
available, INSERT also reports the header-derived offset and the resulting net
offset correction after combining header timing and waveform-based alignment.
In insert mode, a secondary EDF signal is aligned to the primary by applying a constant offset in seconds. If a drift rate is also supplied, or if one has just been estimated, the secondary signal is resampled by cubic spline interpolation to correct the linear time-stretch before insertion. Regions of the primary EDF not covered by the secondary signal are zero-padded.
Parameters
Common parameter:
| Parameter | Example | Description |
|---|---|---|
edf |
edf=secondary.edf |
Secondary EDF to compare against or insert from |
Lag-estimation mode:
| Parameter | Example | Description |
|---|---|---|
pairs |
pairs=C3,C3_ref,C4,C4_ref |
Comma-delimited signal pairs: primary-channel, secondary-channel, repeated |
start |
start=900 |
Window-search start time in seconds; by default INSERT uses a middle-of-recording heuristic |
end |
end=25200 |
Window-search stop time in seconds; by default INSERT uses a middle-of-recording heuristic |
len |
len=300 |
Window length in seconds for local alignment estimation |
inc |
inc=60 |
Increment in seconds between successive windows |
steps |
steps=1000 |
Maximum number of windows to evaluate |
min-peak |
min-peak=0.3 |
Minimum median per-window matched-window correlation required for a window to be accepted |
filt-low |
filt-low=0.5 |
Lower edge of the pre-alignment band-pass filter |
filt-high |
filt-high=15 |
Upper edge of the pre-alignment band-pass filter |
no-filter |
Disable pre-alignment band-pass filtering | |
verbose |
Emit extra per-window and per-pair diagnostics | |
full-search |
Search the full valid lag space for each window; mutually exclusive with offset-range and offset-margin |
|
offset-range |
offset-range=-360,360 |
Explicit absolute search interval in seconds for the secondary-vs-primary offset |
offset-margin |
offset-margin=120 |
Use EDF header start times to estimate the expected offset, then search +/- this many seconds around that value |
auto-try |
Automatically try a short grid of window lengths down to 30 s, keeping start fixed and using inc=len/5 |
|
try-start |
try-start=900,1200 |
Try multiple window start times and keep the best fit |
try-len |
try-len=60,100,180 |
Try multiple window lengths and keep the best fit |
try-inc |
try-inc=5,30,60 |
Try multiple window increments and keep the best fit |
warn-r2 |
warn-r2=0.5 |
Emit a warning if the fitted drift regression R2 falls below this value |
warn-p-ok |
warn-p-ok=0.5 |
Emit a warning if fewer than this fraction of windows pass the quality gate |
warn-peak |
warn-peak=0.35 |
Emit a warning if the median matched-window correlation falls below this value |
no-warn |
Suppress summary alignment-quality warnings |
Insert mode:
| Parameter | Example | Description |
|---|---|---|
insert |
After estimating offset and drift, splice the secondary EDF into the primary EDF in memory | |
sig |
sig=ECG_ref |
Signal(s) from the secondary EDF to insert; default is * |
offset |
offset=-12.5 |
Manual constant shift in seconds for direct insert mode, or manual override of the fitted offset when used with insert |
drift |
drift=-10 |
Manual linear drift term for direct insert mode, or manual override of the fitted drift when used with insert |
secs |
secs=28800 |
Denominator for manual drift, e.g. 28800 for 8 hours |
annot |
annot=MISSING2 |
Reserved for adding missing-data annotations, but not currently implemented |
Output
Estimation mode writes a summary INSERT table plus per-window WIN output. It
also writes a CHS table containing per-pair drift fits. If insert is also
specified, these outputs are written before the in-memory EDF is modified.
Summary INSERT output:
| Variable | Description |
|---|---|
OKAY |
1 if summary alignment quality passed the warning thresholds, else 0 |
N_WIN_ALL |
Total number of windows evaluated before quality gating |
N_WIN |
Number of accepted windows used in the summary fit |
N_OUTLIER |
Number of accepted windows removed as regression outliers |
P_OK |
Fraction of evaluated windows that passed the quality gate |
MEDIAN_SEC |
Median waveform shift in seconds across accepted windows |
MEAN_SEC |
Mean waveform shift in seconds across accepted windows |
MIN_SEC |
Minimum waveform shift in seconds across accepted windows |
MAX_SEC |
Maximum waveform shift in seconds across accepted windows |
RANGE_SEC |
Range of waveform shifts in seconds across accepted windows |
MEDIAN_PEAK |
Median matched-window correlation magnitude across evaluated windows |
MEAN_PEAK |
Mean matched-window correlation magnitude across evaluated windows |
MIN_PEAK |
Minimum matched-window correlation magnitude across evaluated windows |
MAX_PEAK |
Maximum matched-window correlation magnitude across evaluated windows |
INTERCEPT |
Fitted start-of-record waveform shift in seconds |
SLOPE |
Fitted drift slope in seconds per second |
SLOPE_HR |
Fitted drift slope in seconds per hour |
R2 |
R-squared for the fitted waveform-shift-versus-time regression |
IMPLIED_SR |
Implied sample rate of the secondary EDF from the fitted drift |
HDR_OFFSET |
Header-derived offset in seconds from EDF start times alone |
HDR_OFFSET_VALID |
1 if both EDF start times were valid and HDR_OFFSET was available |
TOTAL_OFFSET |
Net offset correction in seconds after combining header timing and waveform-based alignment |
USED_START_SEC |
Window start time used for the selected fit |
USED_LEN_SEC |
Window length used for the selected fit |
USED_INC_SEC |
Window increment used for the selected fit |
AUTO_TUNED |
1 if auto-try or try-* selected the final window settings |
Per-window WIN output:
| Variable | Description |
|---|---|
OK |
1 if the window passed the quality gate, else 0 |
PEAK |
Median matched-window correlation magnitude across signal pairs for that window |
SP |
Estimated waveform shift in sample points for that window |
SEC |
Estimated waveform shift in seconds for that window |
TOT_SEC |
Estimated net offset correction in seconds after combining waveform shift and header offset |
DSEC |
Change in waveform shift relative to the first accepted window |
T1_SEC |
Matched primary time in seconds for that window |
T2_SEC |
Secondary window start time in seconds |
T1_HMS |
Matched primary wall-clock time, when EDF headers are valid |
T2_HMS |
Secondary wall-clock time, when EDF headers are valid |
FIT_USED |
1 if the window entered the drift fit before outlier removal, else 0 |
FIT_OUTLIER |
1 if removed as a drift-fit outlier, 0 if retained, -1 if not fit |
Per-pair CHS output:
| Variable | Description |
|---|---|
INTERCEPT |
Per-pair fitted start-of-record waveform shift in seconds |
SLOPE |
Per-pair fitted drift slope in seconds per second |
SLOPE_HR |
Per-pair fitted drift slope in seconds per hour |
IMPLIED_SR |
Per-pair implied secondary sample rate from the fitted drift |
N_OUTLIER |
Number of outlier windows removed from the per-pair fit |
Direct manual insert mode with only offset=/drift= modifies the in-memory EDF
but does not produce additional estimation tables.
Examples
Estimate offset and drift using the default xcorr-based workflow:
INSERT edf=secondary.edf pairs=C3,C3,C4,C4
Estimate offset and drift using shorter windows and a wider search interval:
INSERT edf=secondary.edf pairs=C3,C3,C4,C4 start=600 len=120 inc=30 offset-range=-360,360
Estimate alignment and then insert signals using the fitted offset and drift:
INSERT edf=secondary.edf pairs=C3,C3,C4,C4 insert sig=ECG,EMG
Insert channels directly once an offset is known:
INSERT edf=secondary.edf sig=ECG,EMG offset=-12.5