Outputs
Commands to rewrite EDFs and signals/annotations in other text-based formats
| Command | Description |
|---|---|
WRITE |
Write a new EDF file |
MATRIX |
Dump signals (and annotations) to a file |
HEAD |
Show a small interval of signal data |
DUMP-RECORDS |
Dump annotations and signals, by EDF record |
RECS |
Dump basic information on EDF record structure |
SEGMENTS |
Dump (discontinuous EDF+) intervals |
SEDF |
Generate a "summary EDF" |
WRITE
Save a (modified) EDF file to disk
Writes a new EDF to disk, that will reflect any manipulation, filtering, or masking, etc, that has been applied.
Parameters
| Parameter | Example | Description |
|---|---|---|
sig |
sig=C3 |
Signal to output (only one) |
edf-dir |
edf-dir=edfs/ |
Set folder where new EDFs should be written |
edf-tag |
edf-tag=v2 |
Add a tag to each new EDF filename |
edf |
edf=f1 |
Write to edf f1.edf |
sample-list |
sample-list=v2.lst |
Name of the new sample-list |
Output
No formal output, other than a message to the log and one or more new EDFs.
Example
To write a set of new EDFs that (for example) have been masked,
filtered and retaining only one signal, given the commands in a file,
say cmd.txt as follows:
EPOCH % Epoch the signals
MASK epoch=1-10 % Set to retain only the first 10 epochs
RESTRUCTURE % Apply the above mask
SIGNALS keep=EEG % Only retain the EEG signal
FILTER bandpass=0.5,4.5 % Apply a bandpass filter to the signal
ripple=0.01
tw=0.5
WRITE edf-dir=newx/ % Write new EDFs, to the folder newx/
edf-tag=v2 % add a 'v2' tag to each EDF
sample-list=newx.lst % create a new sample list pointing to the new EDFs
Running this set of commands:
luna s.lst < cmd.txt
will produce a new folder newx with three new EDFs:
ls newx
learn-nsrr01-v2.edf learn-nsrr02-v2.edf learn-nsrr03-v2.edf
That is, the new EDF filenames have a -v2 tag added. The folder
(which must be specified with a / character in the edf-dir
argument) will be created if it does not exist. In addition, Luna
creates a new sample-list called
newx.lst, that points to these new EDFs:
cat newx.lst
nsrr01 newx/learn-nsrr01-v2.edf
nsrr02 newx/learn-nsrr02-v2.edf
nsrr03 newx/learn-nsrr03-v2.edf
Note that Luna appends each item to this list, and so you may want to delete it before running the command, if it already exists.
We can use the new sample list to check the properties of the new set of EDFs:
luna newx.lst -s DESC
As expected, we do in fact see that the EDFs are now only 10 epochs in length and contain only a single channel: (here, showing output only for the first EDF):
EDF filename : newx/learn-nsrr01-v2.edf
ID : nsrr01
Clock time : 21:58:17 - 22:03:17
Duration : 00:05:00
# signals : 1
# EDF annotations : 1
Signals : EEG[125]
Finally, to check the filtering, we can use the MATRIX command to
dump the raw signals to a file. First from the original EDF (for the first 10 epochs only):
luna s.lst 1 -s "MASK epoch=1-10 & RE & MATRIX sig=EEG file=old.txt"
luna newx.lst 1 -s "MATRIX file=new.txt"
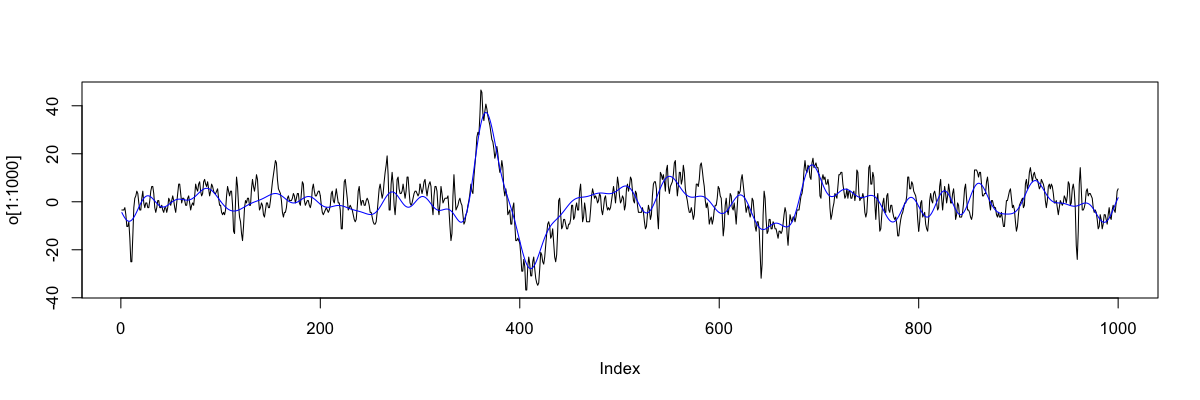
In R
o <- scan("old.txt",skip=1)
n <- scan("new.txt",skip=1)
plot( o[1:1000] , type="l" )
lines( n[1:1000] , col="blue" )
MATRIX
Dumps signal information to a file
For one or more signals of similar sampling rates, this command generates a text file containing the raw signal data.
Parameters
| Parameter | Example | Description |
|---|---|---|
file |
file=signals.txt |
Required parameter, to specify the filename for the output |
sig |
sig=C3,C4 |
Restrict output to these signals/channels |
hms |
hms |
Add a clock-time column in hh:mm:ss format |
hms2 |
hms2 |
Add a clock-time column in hh:mm:ss:microsecond format |
annot |
annot=X,Y |
Add columns with values 1/0 to indicate the presence/absence of that annotation |
min |
min |
Minimal output to show only signal information (no headers or lead columns) |
Output
All output is written to a text file as specified by the file parameter. For example:
ID E S SP T EOG-L EOG-R EMG EEG
id01 1 0 0 0 -6.07448 4.18193 31.044 0.763126
id01 1 0 1 0.00390625 -0.030525 2.83883 52.7778 11.5079
id01 1 0 2 0.0078125 7.23443 2.22833 45.3297 23.0464
id01 1 0 3 0.0117188 11.569 2.10623 23.4127 29.4567
id01 1 0 4 0.015625 12.9731 2.16728 8.82173 31.2882
id01 1 0 5 0.0195312 12.79 2.22833 -4.73138 30.7387
id01 1 0 6 0.0234375 12.1795 2.28938 -13.6447 29.6398
id01 1 0 7 0.0273438 11.6911 2.22833 -14.1331 28.663
id01 1 0 8 0.03125 11.3858 2.22833 -11.8742 28.0525
id01 1 0 9 0.0351562 11.2027 2.16728 -13.4615 27.6862
... cont'd ...
Here, the first five columns are:
ID: individual/EDF IDE: epoch numberS: elapsed time in seconds (integer)SP: sample point in the EDF recordT: elapsed time in seconds
The subsequent columns represent the channels in the EDF (or those
specified by the sig parameter).
If the min (or minimal) parameter is specified, then the header
and the first five columns are omitted.
If the hms parameter is specified, then an additional column HMS
is added, which is the clock-time in hh:mm:ss format. If hms2 is
specified instead of hms, this field is printed with micro-second
resolution.
If the annot parameter is specified, additional columns are added
with the same names as the annotations specified, e.g. annot=X,Y
will add two columns X and Y. For each sample point, these
columns will have a 0 or 1 value to indicate whether or not that
annotation was present at that point.
HEAD
Writes a small amount of signal data to standard output
This command is useful to sanity-check signals and epochs, by outputting just a small amount of data (e.g. a few seconds worth) for one or more channels, for a given epoch.
If multiple channels are output, they must all have the same sample rate.
Parameters
| Parameter | Example | Description |
|---|---|---|
sig |
C3,C4 |
Show these channels (default: all channels) |
epoch |
100 |
Show epoch 100 (default: first epoch) |
sec |
0.5 |
Only show a fixed duration (secs) of the epoch (default: whole epoch) |
Output
All output is sent to the console (stdout) and so can be redirected,
etc. Output is tab-delimited; the first three columns are T
(elapsed seconds from start of EDF), SEC (elapsed seconds in this
segment of data) and SP (sample point in this segment of data,
starting at 0). The subsequent columns are for the requested channels.
Example
To output 0.05 second's worth of data for two EOG channels, from a particular epoch of an EDF (here, 222):
luna s.lst 1 -s HEAD epoch=222 sig=EOG_L,EOG_R sec=0.05 > o.txt
T SEC SP EOG_L EOG_R
6630 0 0 -8.21123 -6.37973
6630 0.00390625 1 -7.72283 -5.52503
6630.01 0.0078125 2 -6.68498 -4.30403
6630.01 0.0117188 3 -5.76923 -3.20513
6630.02 0.015625 4 -4.91453 -2.71673
6630.02 0.0195312 5 -3.26618 -2.47253
6630.02 0.0234375 6 -0.457875 -1.98413
6630.03 0.0273438 7 2.28938 -1.31258
6630.03 0.03125 8 3.93773 -0.763126
6630.04 0.0351562 9 4.30403 -0.0915751
6630.04 0.0390625 10 4.18193 0.763126
6630.04 0.0429688 11 3.99878 1.43468
6630.05 0.046875 12 3.87668 1.67888
Note
Note that T may have limited numerical precision in the output.
This command is intended for quick reviews of signals (i.e. to see
units, scales, etc).
DUMP-RECORDS
Writes detailed annotation and signal data to standard output
This command is unlikely to be of great utility to most users. It
dumps detailed information about the signals and annotations to
stdout.
Parameters
| Parameter | Description |
|---|---|
no-signals |
Do not show signal data |
no-annots |
Do not show annotation information |
Output
All output is sent to the console (stdout) and so can be redirected,
etc. Output is organized by EDF record, e.g.:
Record 1 of 40920 total (30 retained)
Generic Annotations-----------------------
wake wake 0.00->30.00 wake[flag]=.
EDF Annotations--------------------------
Signal 15 EDF Annotations
<0||(time-stamp, secs)>
SaO2 has a sample rate of 1 Hz, and so there is only one entry per record:
s = 0
interval = 0-999999999999
RECORD-DUMP SaO2 rec=0 1/1 0 0 95.115587
PR:
s = 1
interval = 0-999999999999
RECORD-DUMP PR rec=0 1/1 0 0 74.222934
s=2) is EEG(sec), which has a sample rate of 125Hz (and so 125 entries here):
s = 2
interval = 0-999999999999
RECORD-DUMP EEG(sec) rec=0 1/125 0 0 -0.49019608
RECORD-DUMP EEG(sec) rec=0 2/125 8000000000 0.008 1.4705882
RECORD-DUMP EEG(sec) rec=0 3/125 16000000000 0.016 6.372549
RECORD-DUMP EEG(sec) rec=0 4/125 24000000000 0.024 3.4313725
RECORD-DUMP EEG(sec) rec=0 5/125 32000000000 0.032 -10.294118
RECORD-DUMP EEG(sec) rec=0 6/125 40000000000 0.04 -8.3333333
RECORD-DUMP EEG(sec) rec=0 7/125 48000000000 0.048 1.4705882
RECORD-DUMP EEG(sec) rec=0 8/125 56000000000 0.056 -0.49019608
RECORD-DUMP EEG(sec) rec=0 9/125 64000000000 0.064 -0.49019608
RECORD-DUMP EEG(sec) rec=0 10/125 72000000000 0.072 7.3529412
RECORD-DUMP EEG(sec) rec=0 11/125 80000000000 0.08 11.27451
... cont'd ...
- a standard
RECORD-DUMP - signal name
- record number (starting at 0)
- the sample point per record (e.g.
1/125,2/125, etc) - the starting time for that sample point in time units
- time in seconds
- the value of the signal
For most purposes, the MATRIX command will likely be an
easier route to achieve direct access to the signals in an EDF. (In
fact, this command was in large part only added to assist in debugging
during Luna development, but is described here for completeness.)
RECS
Dumps basic information about record structure to standard output
Parameters
None.
Output
Text written to the log/console; this command does not generate any output through Luna's standard output mechanism.
Example
Taking the first tutorial EDF (where EDF is one second):
luna s.lst 1 -s SUMMARY
Rec. dur. (s) : 1
we extract a subset of the data (epochs 5 through 8, defaulting to 30-second epochs) and then call the RECS command:
luna s.lst 1 -s 'MASK epoch=5-8 & RECS'
The output (to the console) is as follows (abridged):
RECS nsrr01 1 121 120/40920 120.00->121.00 5;120.00->150.00
RECS nsrr01 2 122 120/40920 121.00->122.00 5;120.00->150.00
RECS nsrr01 3 123 120/40920 122.00->123.00 5;120.00->150.00
RECS nsrr01 4 124 120/40920 123.00->124.00 5;120.00->150.00
RECS nsrr01 5 125 120/40920 124.00->125.00 5;120.00->150.00
...
RECS nsrr01 115 235 120/40920 234.00->235.00 8;210.00->240.00
RECS nsrr01 116 236 120/40920 235.00->236.00 8;210.00->240.00
RECS nsrr01 117 237 120/40920 236.00->237.00 8;210.00->240.00
RECS nsrr01 118 238 120/40920 237.00->238.00 8;210.00->240.00
RECS nsrr01 119 239 120/40920 238.00->239.00 8;210.00->240.00
RECS nsrr01 120 240 120/40920 239.00->240.00 8;210.00->240.00
The columns are:
RECScommand label- current EDF ID
- record number with respect to current in-memory EDF representation (always starts 1)
- record number with respect to original EDF file (starting 121, i.e. the start of the fifth epoch)
- number of records in the current in-memory EDF / total number of records in the on-file EDF
- start and stop of this record (in seconds)
- (for epoched data) the epoch number (e.g. 5-8) that record belongs to, and the epoch start/stop (seconds)
Four 30-second epochs (5,6,7 and 8) implies 120 seconds; the start/stop times correspond to this, as expected: 240-121+1 = 120.
SEGMENTS
Reports the number of contiguous segments in an EDF/EDF+
Identifies and reports on the contiguous segments in a file; this is
primarily of use for EDF+ files with discontinuous signal data.
Internally, Luna represents signal data as equivalent to an EDF+ after
any masks that remove epochs have been applied;
therefore, the SEGMENTS command can also be used in this
context too.
Parameters
| Parameter | Description |
|---|---|
annot |
Add annotations to mark segment/gap boundaries |
Output
Basic EDF header information (strata: none)
| Variable | Description |
|---|---|
NR |
Number of segments |
Per-segment information (strata: SEG)
| Variable | Description |
|---|---|
START |
Segment start (seconds) |
STOP |
Segment stop (seconds) |
START_HMS |
Segment start (hh:mm:ss) |
STOP_HMS |
Segment stop (hh:mm:ss) |
DUR_HR |
Segment duration (hours) |
DUR_MIN |
Segment duration (minutes) |
DUR_SEC |
Segment duration (seconds) |
Example
Taking the first tutorial EDF, which is a continuous EDF file:
luna s.lst 1 -s SEGMENTS
We'd expect just a single segement to be reported, and this is indeed what we see:
destrat out.db +SEGMENTS
ID NSEGS
nsrr01 1
destrat out.db +SEGMENTS -r SEG
ID SEG DUR_HR DUR_MIN DUR_SEC START START_HMS STOP STOP_HMS
nsrr01 1 11.3667 682 40920 0 21.58.17 40920 09.20.17
In contrast, if we break up the data (e.g. using the
MASK command, then the data are implicitly
represnted as a discontinuous EDF+ after restructuring:
luna s.lst 1 -t out -s 'MASK epoch=9-12,22-23,70-74 & RE & SEGMENTS'
Hint
Note, purely to illustrate different options, here we used Luna's -t output option rather than -o
Now the SEGMENT command shows the expected three segments:
cat out/nsrr01/SEGMENTS.txt
ID NSEGS
nsrr01 3
cat out/nsrr01/SEGMENTS-SEG.txt
ID SEG DUR_HR DUR_MIN DUR_SEC START START_HMS STOP STOP_HMS
nsrr01 1 0.0333333 2 120 240 22.02.17 360 22.04.17
nsrr01 2 0.0166667 1 60 630 22.08.47 690 22.09.47
nsrr01 3 0.0416667 2.5 150 2070 22.32.47 2220 22.35.17
If annotations are added, the default labels are segment and gap (with the instance ID
reflecting the number of each). These labels can be changed with the special variables annot-segment and annot-gap (i.e.
if there is a pre-existing annotation with that label:
luna s.lst annot-gap=G -s 'SEGMENTS annot & WRITE-ANNOTS file=^.annot'
See this vignette for an example of using SEGMENTS annot to track changes from an EDF+D to an EDF file.
SEDF
Generates a summary-EDF and writes it to disk
For data channels only (i.e. not any EDF+ annotation channels), the SEDF command
creates a "summary" EDF
| Option | Description |
|---|---|
sig |
Signals to output (if not given, all signals) |
sedf-dir |
Directory for SEDFs (if not working directory) |
This creates an EDF in which contains epoch-level summary statistics of the original EDF. That is, if the original EDF has 1000 30-second epochs, the SEDF will comprise only 1000 records, where each record has a duration of 30 seconds, and contains only a single value. For a given channel, e.g. CZ, the SEDF will contain three new channels, depending on the type of that channel. For a channel called XYZ in the original:
- for oscillatory signals (e.g. EEG, EMG, etc), the three Hjorth parameters:
XYZ_H1,XYZ_H2andXYZ_H3 - for other signals (e.g. light or heart rate, etc), the mean, min and max:
XYZ_M,XYZ_L,XYZ_U
The idea is that this SEDF file is an effective thumbnail for the EDF, which can be quickly loaded and rendered, e.g. by a viewer application.